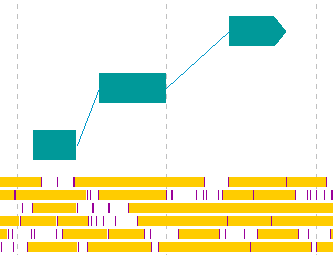
- CDS candidates can be registered as new features from the stop lane stop absence region of the frame lane.
- This is not a CDS prediction function. The GT-AG rule is ignored.
Operation
- We display the feature lane and frame lane in the main feature map.
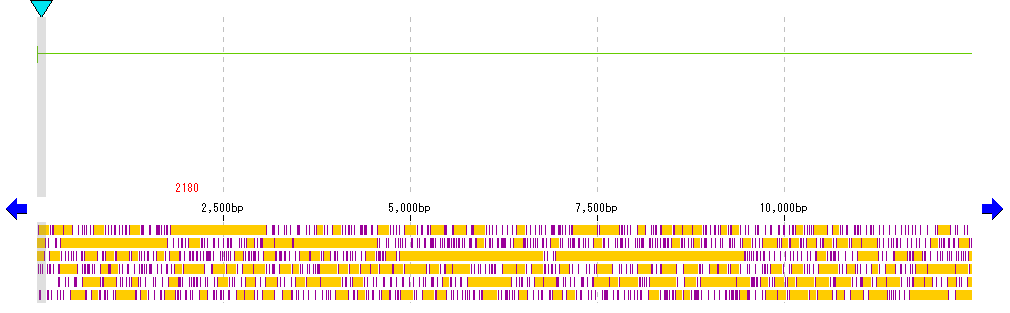
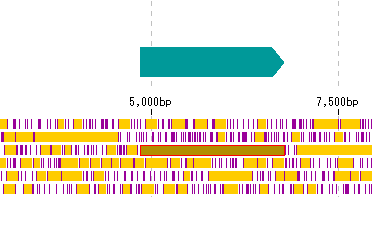
- Right-click on the stop lane stop codon absent area (orange area in the above figure) in the frame lane.
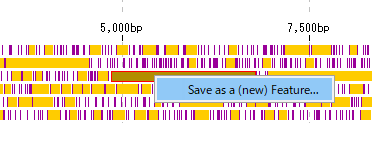
- The description window will be displayed.
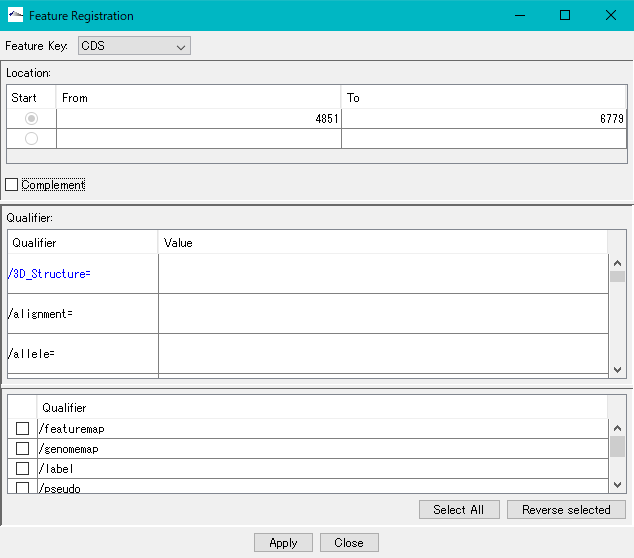
- Click "Apply".
- The CDS corresponding to the selected area is newly registered.
- If CDS display is set in the feature lane, it will be displayed as follows.

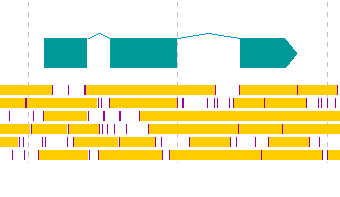
- Register CDS candidates consisting of multiple exons in eukaryotic organisms.
- Display the frame lane in the main feature map.
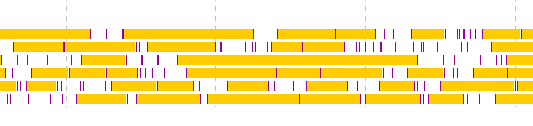
- Click from the first exon candidate to the region immediately before the last exon.
- If you click the last exon candidate before right-clicking the mouse, it will be out of the selection.
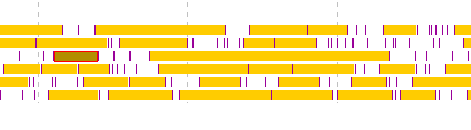
- Right click on the final exon candidate.
- A menu will be displayed and click on "Save as a (new) Feature".
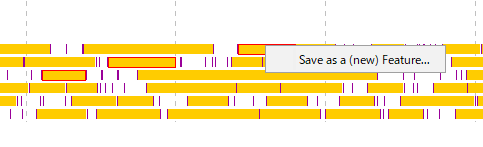
- The description window will be displayed.
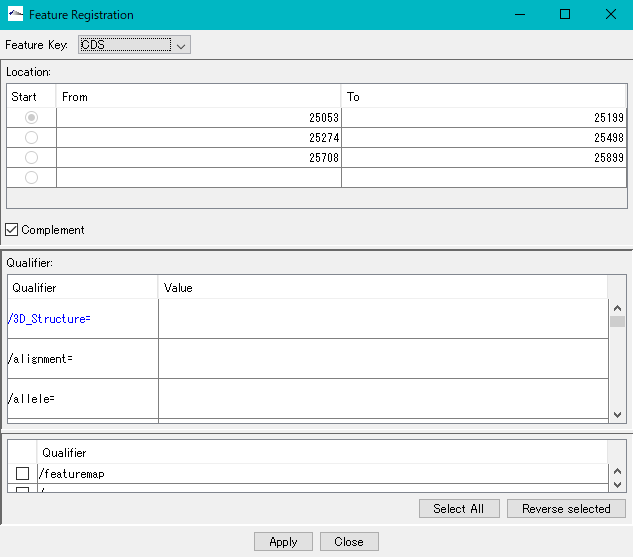
- Position information of the number of areas specified for Location is set.
- Click "Apply".
- A multi-exon CDS will be displayed on the feature lane.

- It is also possible to draw for each frame to which each exon belongs.