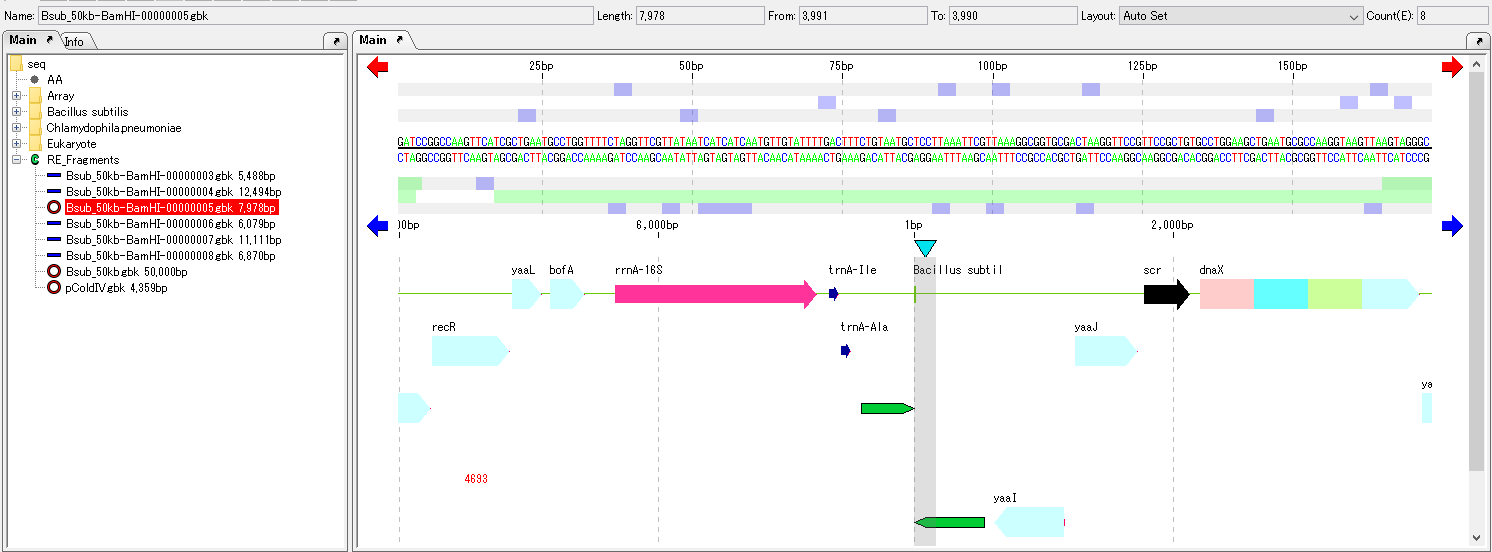
Ligate the ends of one DNA sequence.
This is called self-ligation.
When self-ligation is performed in IMC, its DNA sequence attribute is changed from Linear to Circular.
Therefore, if you display that feature map, you can scroll horizontally indefinitely in the same direction if you could only scroll to the end until then.
In order to return this to the original linear DNA, it is necessary to cut at the same position.
If both ends are cleaved with the same restriction enzyme, digestion with the restriction enzyme cleaves it back to the original linear DNA.
Operation
- Load the base sequence file that self-ligates into the main feature map.
- From the menu, choose Cloning -> Simple Ligation.
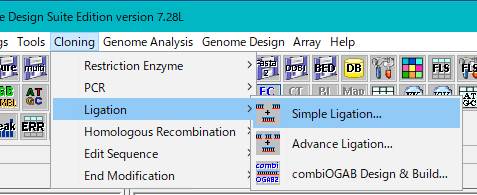
- The Ligation dialog will be displayed.
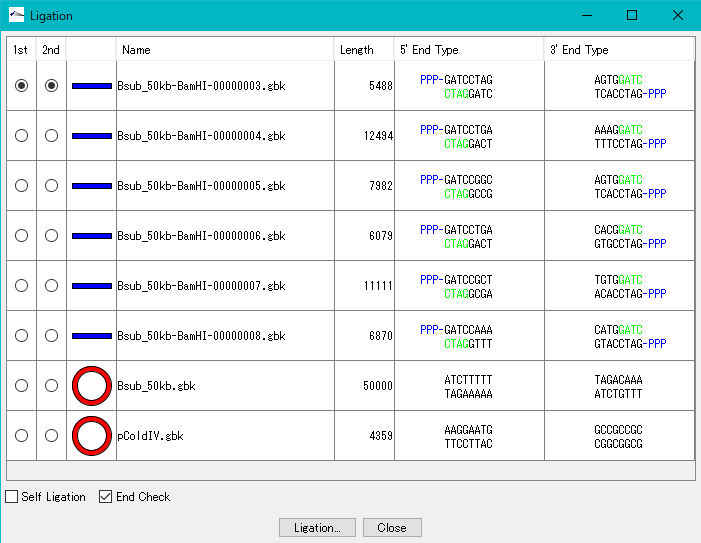
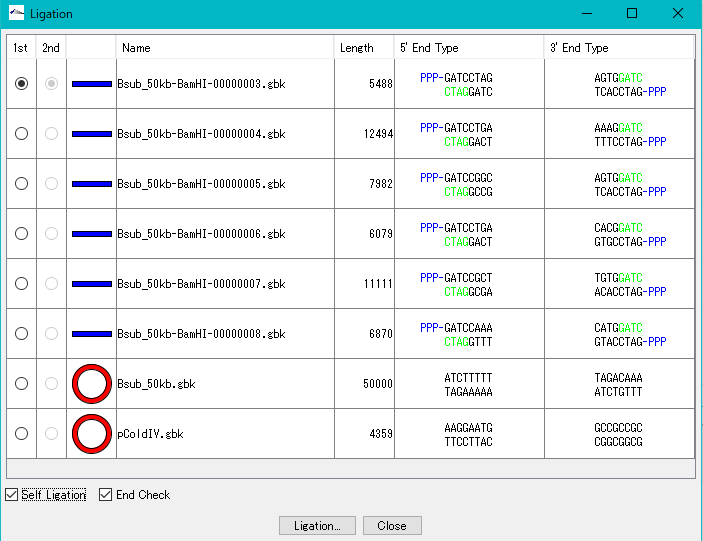
- In the list, all the sequences loaded in the main current directory are displayed.
- Check the Self Ligation check box.
- 2nd radio button becomes Inactive.
- Turn on the 1st radio button of the self-ligation sequence.
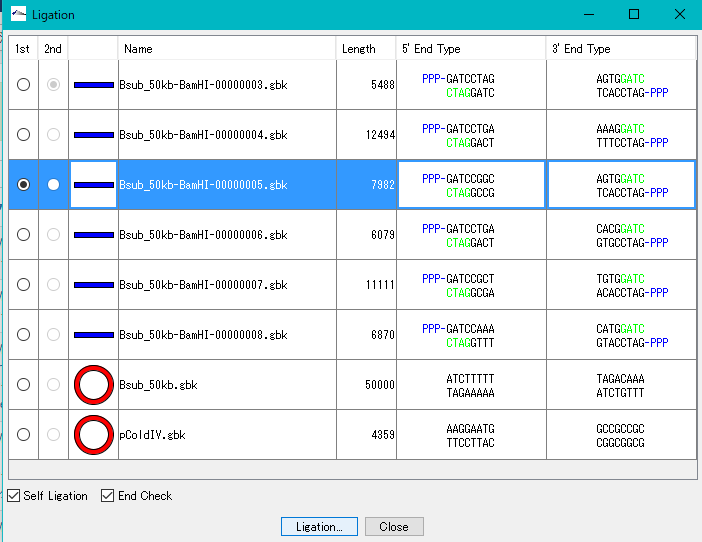
- Click "Ligation".
- A confirmation message will be displayed.
- Click "Yes (Y)".
- Ligation is performed,
- A completion message is displayed.
- Click OK.
- The DNA shape of the target sequence in the Ligation dialog becomes a
 mark.
mark. 
- Click "Close".
- The Ligation dialog closes.
- The DNA shape of the corresponding sequence in the main current directory is also a
 mark.
mark. 
- We will make that array the current sequence.
- A map of the self-ligated sequence is displayed in the main feature map on the right.
- Scroll in both directions, you can scroll indefinitely and display the ligated base position in the center.