GT 11A Import BAM Files using Training Data
In order to get used to the operation of GT, we will use BAM import mechanism and operation using training data.
This BAM file is the assembly result of NGS lead by Velvet.
Operation
Start GT.
In the initial state, the main window is displayed as follows.
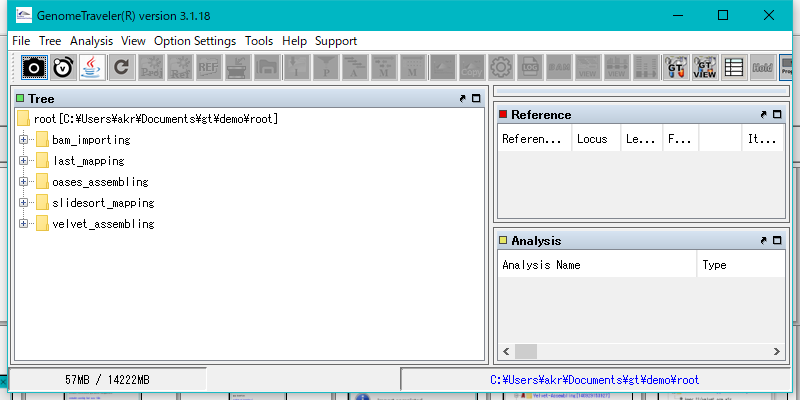
Right-click on the Root node in the TREE pane.
The menu will be displayed.
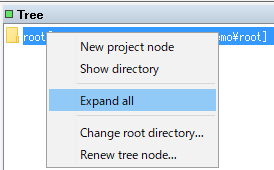
Select Expand All.
All nodes in the TREE pane are expanded.
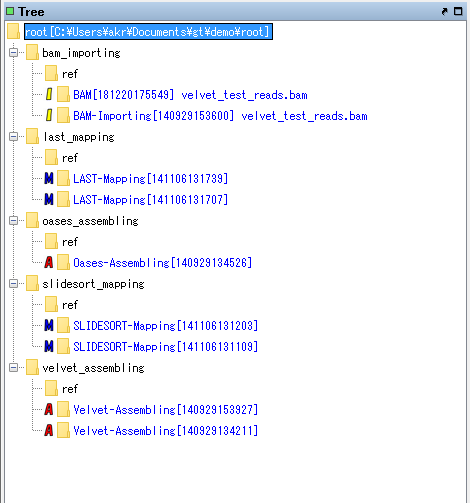
Use the top "bam_importing".
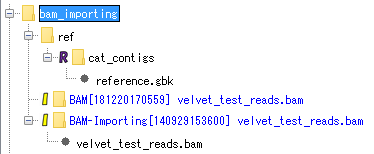
Right click on the "bam_importing" node.
The menu will be displayed.
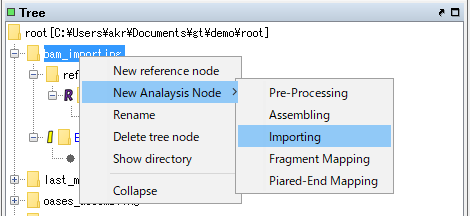
Select New Analysis Node -> Importing.
The BAM - Importing dialog will be displayed.
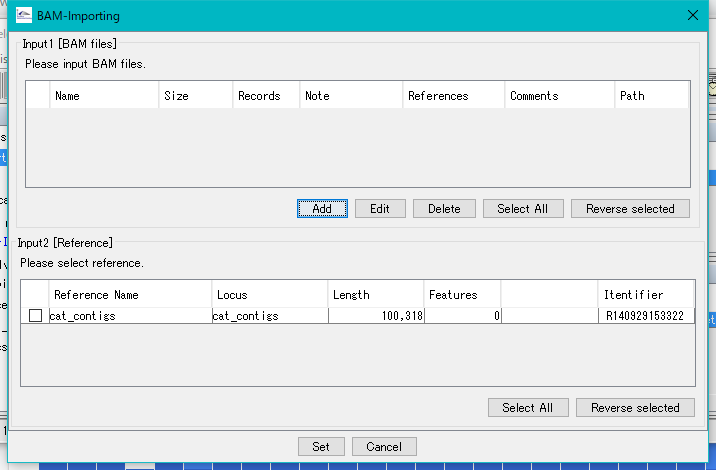
Click Add.
The "Add BAM Files (s)" dialog is displayed.
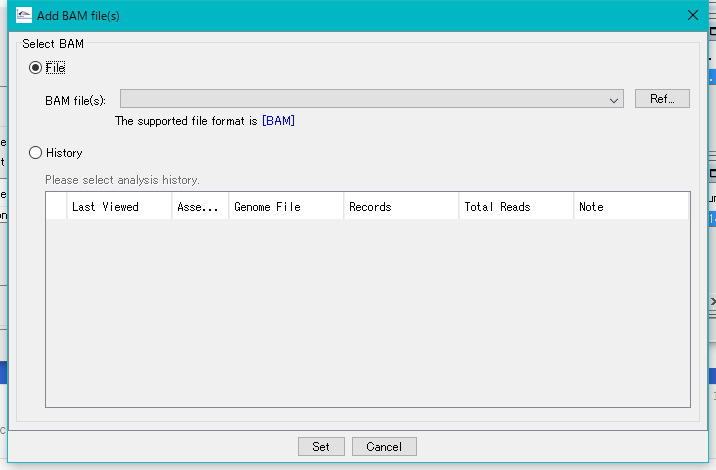
Click Ref ... and select the BAM file in the demo data area.
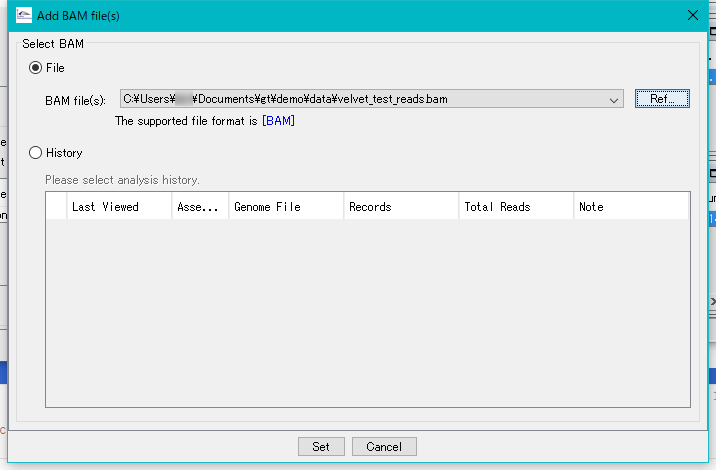
Click "Set".
Return to the BAM - Importing dialog.
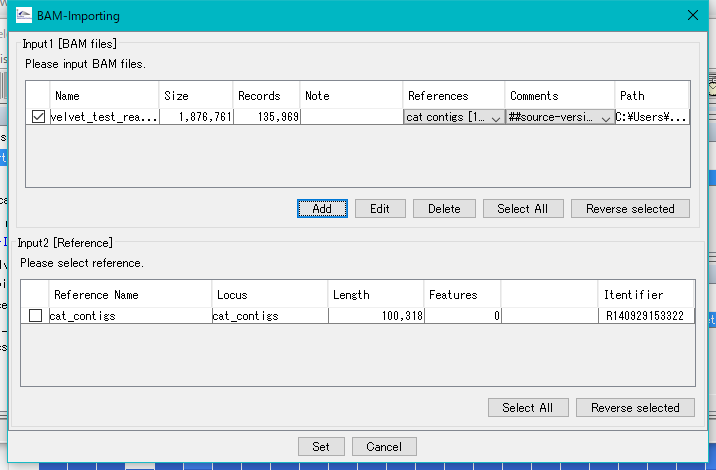
Check all check boxes.
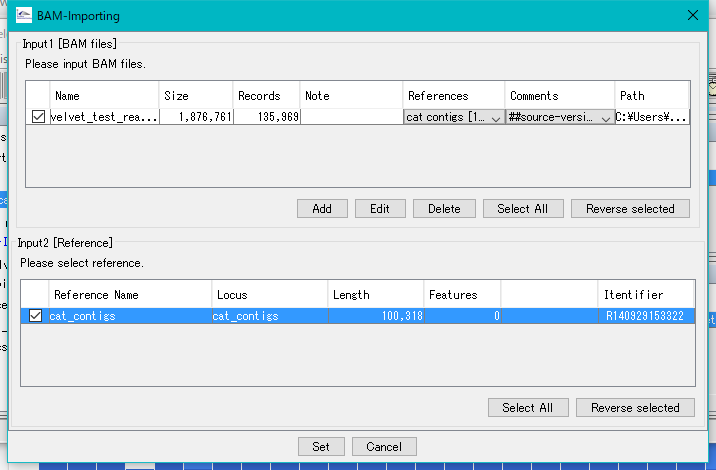
Click Set.
A Start Importing confirmation message is displayed.
Click "Yes".
Execution of the import starts and a progress message is displayed.
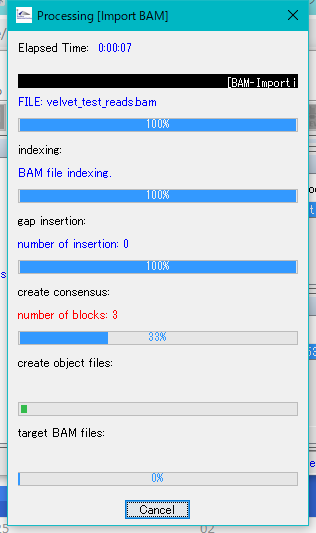
When the import is completed, a completion message is displayed.
Bam_importing node in the TREE pane A new import node is created.
Click on the new node to expand it.Right click mouse over Velvet node.
The menu will be displayed.
Select "Show BAM Information".
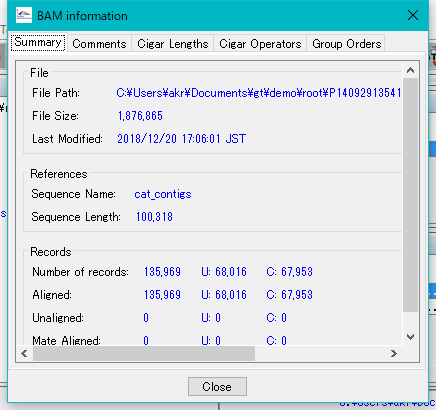
The BAM Information dialog will be displayed.
Click the tab to switch the panes.
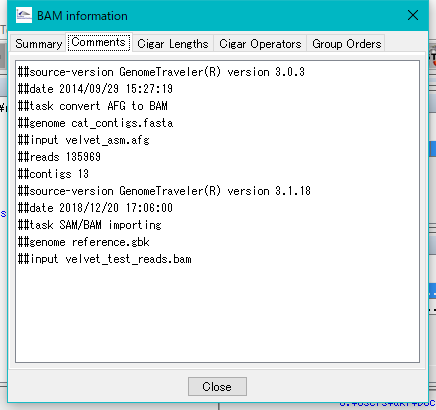
Right click mouse over Velvet node.
The menu will be displayed.
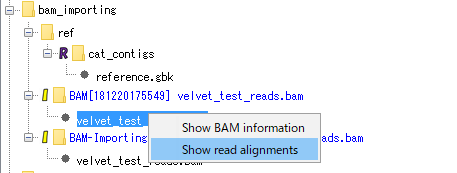
Select "Show Read Alignment". The Alignment Viewer is displayed.
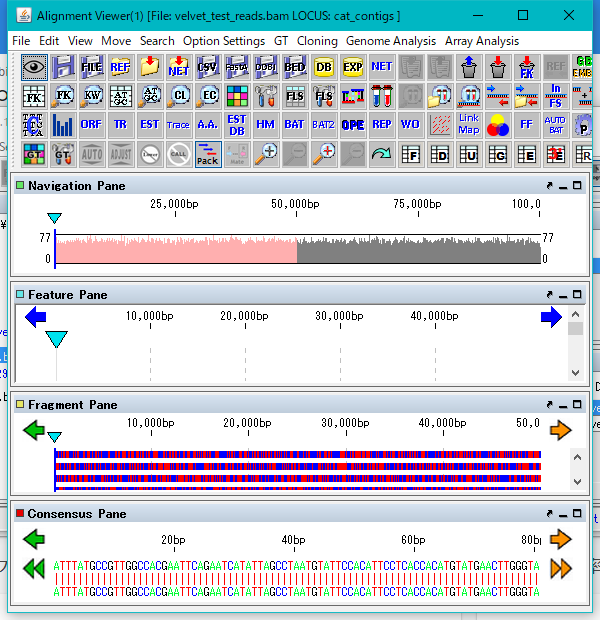
The Alignment Viewer can be resized.
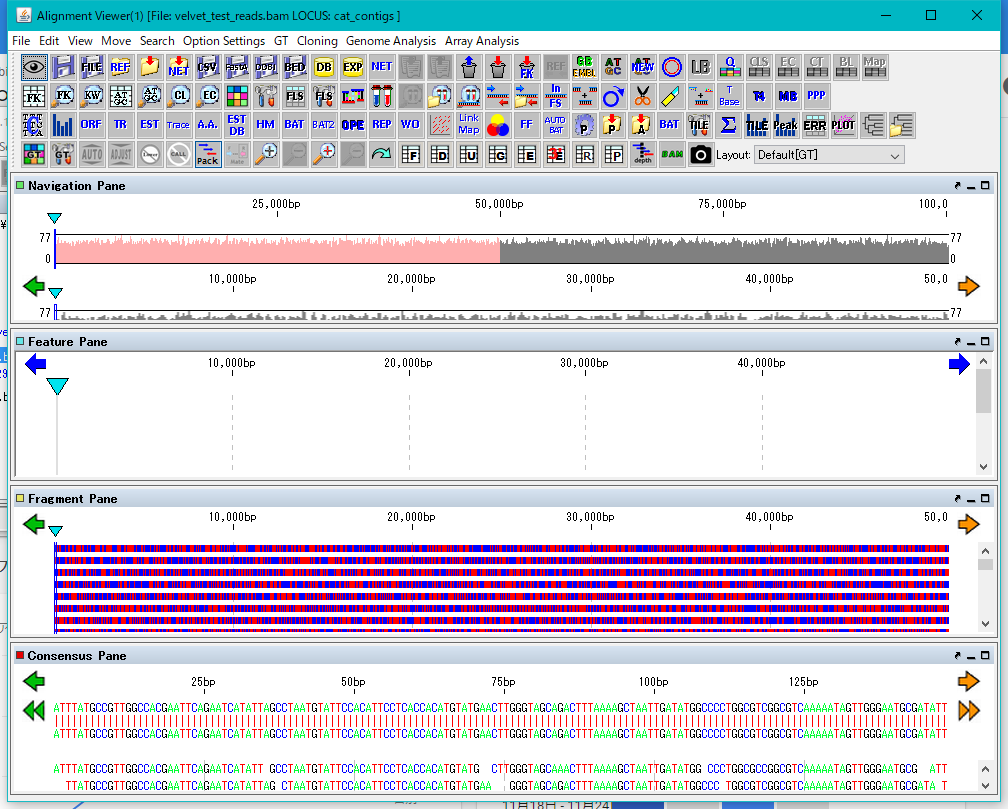
 Dongle License (HW Key)
Dongle License (HW Key) Feature Map
Feature Map Management and Operations of Feature Keys
Management and Operations of Feature Keys Sequence and Data Input and Output
Sequence and Data Input and Output GenBank EMBL Viewer
GenBank EMBL Viewer Sequence Viewer
Sequence Viewer Annotation Viewer
Annotation Viewer Circular Genome Viewer-Designer
Circular Genome Viewer-Designer Plasmid Map Viewer-Designer
Plasmid Map Viewer-Designer Trace Viewer - Editor
Trace Viewer - Editor Phylogenetic Tree Viewer
Phylogenetic Tree Viewer Feature Key Search
Feature Key Search Keyword Search
Keyword Search Pattern Search
Pattern Search Priming Site Search
Priming Site Search Batch Homology Search
Batch Homology Search Restriction Enzyme
Restriction Enzyme Primer Design
Primer Design PCR Reaction
PCR Reaction Ligation
Ligation Fragment Modification
Fragment Modification DNA Content Analysis
DNA Content Analysis Codon Analysis
Codon Analysis ORF Analysis
ORF Analysis Database Management
Database Management Multiple Circular Genome Map
Multiple Circular Genome Map Dot Plot Analysis
Dot Plot Analysis Venn Diagram Analysis
Venn Diagram Analysis Reverse Complement
Reverse Complement Settings
Settings