IMC Version 8.135 Release Note
The latest version of IMC, version 8.135, has been released.
- MacOS 10.15, 11, 12, 13, 14
- Windows 10, 11
Fixes and changes
- Auto update function is restored.
IMC Version 8.133 Release Note
The latest version of IMC, version 8.133, has been released.
- MacOS 10.15, 11, 12, 13, 14
- Windows 10, 11
Bug
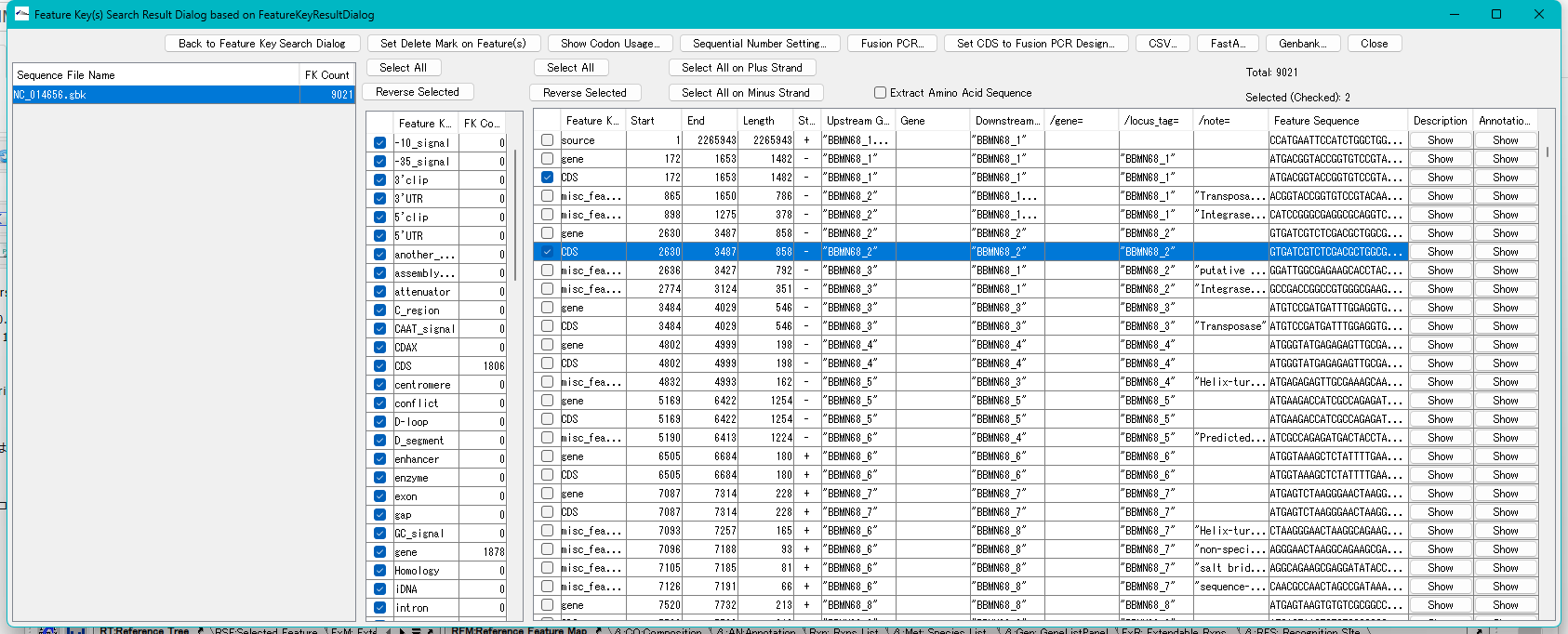
- There was a bug in the Feature Key Search Result Dialog where the button positions and display were distorted.
Fixes and changes
The display positions of the buttons were overlapping, and some buttons were not displayed, some buttons were out of place, and some buttons had functions that were difficult to understand, so we have improved the button names and display positions.
Feature Key Search Result Dialog Overall Operation Buttons
- Button label change
- Back -> Back to Feature Key Search Dialog
- The position to return to has been clarified by pressing Back.
- Back -> Back to Feature Key Search Dialog
- Delete -> Set Delete Mark on Feature(s)
- The Delete button does not immediately delete the selected feature, but simply adds a deletion mark to that feature.
- For this reason, it has been clarified that it means to add a deletion mark.
- The Delete button does not immediately delete the selected feature, but simply adds a deletion mark to that feature.
Feature Key Table Operation Buttons
- Select All: The display position overlapped with the Select All button in the Feature Key Detail Table. Displayed at the top of the Feature Key Table.
- Reverse Selected: Moved to the top of the Feature Key Table.
Feature Key Detail Table Operation Buttons
- Select All: Selects all features in the Detail Table.
- Reverse Selected: Reverses the current selection in the Detail Table.
- Select All on Plus Strand: Selects all features on the plus strand of.
- Select All on Minus Strand: Selects all features on the plus strand of.
Feature Key Detail Table Check Boxes
- Extract Amino Acid Sequence: Extracts CDS features as amino acid sequences.
Feature Key Detail Table Labels
- Total: Total number of features with the currently selected feature key
- Selected (Checked): Total number of features currently selected in the Feature Key Detail Table
IMC Version 8.132 Release Note
The latest version of IMC, version 8.132, has been released.
IMC Version 8.131 The release of 8.131 has been cancelled due to a problem with the installer.
- MacOS 10.15, 11, 12, 13, 14
- Windows 10, 11
Bug
- In the Legacy Primer List dialog, there was a bug where the results of new primer registrations were not reflected immediately.
Fix
- The above bug has been fixed.
Specification change
- It is now possible to set a setting to stop the automatic registration of links to genome sequence files that have the priming site of a primer in a primer record.
IMC Version 8.130 Release Note
The latest version of IMC, version 8.130, has been released.
MacOS 10.15, 11, 12, 13, 14
Windows 10, 11
Bug
- There was a bug in the settings of the library file related to Excel operations.
- Some menu items from the TREE node of NGS Workbench (formerly GenomeTraveler) did not pop up depending on the situation.
Fix
- The above bugs have been fixed.
IMC Version 8.125 Release Note
The latest version of IMC, version 8.125, has been released.
The release of version 8.124 has been skipped.
Supported OS
- MacOS 10.14 and 10.14 are no longer supported.
Bug fixes
- Fixed IMCPH initialization problem.
IMC Version 8.126 Release Note
The latest version of IMC, version 8.126, has been released.
- MacOS 10.14 and 10.14 are no longer supported.
- MacOS 10.15, 11, 12, 13, 14
- Windows 10, 11
Bug fixes
- Fixed bugs in Primer Manager.
IMC Version 8.129 Release Note
The latest version of IMC, version 8.128, has been released.
Supported OS
- Windows: 10, 11
- MacOS: X.15, 11, 12, 13, 14
Bug
- There was a bug where the cloning function button tools (multiple buttons, such as the restriction enzyme button) in the toolbox were grayed out and could not be clicked.
Fix
- The above bug has been fixed.
IMC Version 8.128 Release Note
The latest version of IMC, version 8.128, has been released.
- MacOS 10.15, 11, 12, 13, 14
- Windows 10, 11
Bug fixes
Fixed a bug that caused Show License (formerly About IMC) to not display.
Specification changes
The old About IMC menu item has been changed to Show License.
 Dongle License (HW Key)
Dongle License (HW Key) Feature Map
Feature Map Management and Operations of Feature Keys
Management and Operations of Feature Keys Sequence and Data Input and Output
Sequence and Data Input and Output GenBank EMBL Viewer
GenBank EMBL Viewer Sequence Viewer
Sequence Viewer Annotation Viewer
Annotation Viewer Circular Genome Viewer-Designer
Circular Genome Viewer-Designer Plasmid Map Viewer-Designer
Plasmid Map Viewer-Designer Trace Viewer - Editor
Trace Viewer - Editor Phylogenetic Tree Viewer
Phylogenetic Tree Viewer Feature Key Search
Feature Key Search Keyword Search
Keyword Search Pattern Search
Pattern Search Priming Site Search
Priming Site Search Batch Homology Search
Batch Homology Search Restriction Enzyme
Restriction Enzyme Primer Design
Primer Design PCR Reaction
PCR Reaction Ligation
Ligation Fragment Modification
Fragment Modification DNA Content Analysis
DNA Content Analysis Codon Analysis
Codon Analysis ORF Analysis
ORF Analysis Database Management
Database Management Multiple Circular Genome Map
Multiple Circular Genome Map Dot Plot Analysis
Dot Plot Analysis Venn Diagram Analysis
Venn Diagram Analysis Reverse Complement
Reverse Complement Settings
Settings