Feature Key and Feature
Subcategories
Management and Operations of Feature Keys 2
Default feature key
Create, edit, delete feature keys
Template function
Feature key search
Attributes of Feature Key 0
Feature Keys
Attributes of Feature Keys
IMC Original Set Feature Keys 1
IMC has its original feature key set.
These Original feature keys exist to control the drawing and editing of features.
There is also a function to remove all original feature keys at once.
Types and Roles of Qualifier 0
Types and Roles of Qualifiers are explained here.
View and Edit Qualifier 4
It collects information on the display method and editing method of Qualifier showing the attribute of Feature.
The following shows two ways of displaying the value of Qualfier described in Feature.
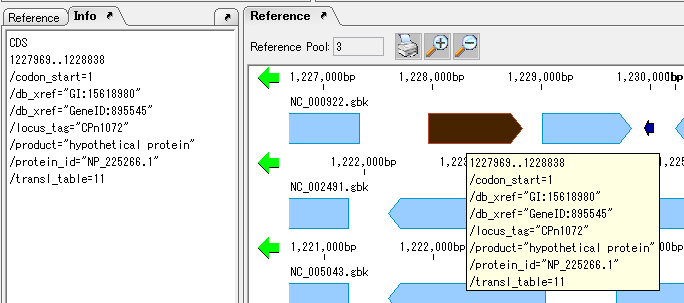
Feature Position on Genome Sequence 1
Explanation on the location of the feature on the genome sequence.
Feature Fragmentation 0
Fragmentation of features by restriction enzymes and PCR amplification.
Feature Synthesis 0
Explain synthesis of features by PCR or ligation.
Feature Fusion 1
Feature Fusion is a feature that merges features belonging to the same Feature Key in two identical positions into one. Qualifiers recorded in each feature to be merged are kept equal on either side as the Qualifier of the merged feature.
This feature makes it easy to combine multiple annotations separately attached to the same feature into one.
Feature Operators 1
The Feature operators are logical arithmetic functions of performing logical operations on two mutually overlapping features belonging to the same Feature Key and generating new features.
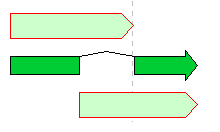
A feature where the upper and lower arrows in the above figure overlap, and the center divided arrow is a computed feature generated.
Link and Refer on Feature 1
You can link to individual features and activate the visualization tool as follows.
- Molecular structure files and its visualization tool,
- Document file (pdf) and PDF display tool
Feature Mapping 0
Mapping to Individual Features
Feature Export 3
You can select the base sequence or amino acid sequence of the feature and export it as a file.
The Feature Export function has the following.
- Export Feature (CSV) ... -> Export features and sequences in CSV format.
- Export Feature & Sequence (DDBJ) ... exports features and sequences in DDBJ format.
- Export Feature (BED) ... -> Export features in BED format.
Besides, using the function to output the result of the search function as a file actually becomes a feature export.
- Export from Feature Key Search result dialog
- Export from Keyword Search result dialog
- Export from Classification Search results dialog
Attach Sequential Number to Feature (Qualifier Editing) 3
Add a serial number to any Feature Qualifier.
You can select Qualifier to be serial number.
Normally, users can not register their original qualifiers, but you can create your own original qualifier that is used only when adding sequential numbers.
Sequential numbers consist of prefixes + sequence numbers, and consecutive numbers are further generated with a starting number and an increment.
The features that are subject to the sequential numbering are added in order from the one closest to the base position of the genome base position.
Not only features belonging to one type of feature key, but also features belonging to multiple feature keys can be mixed and added sequentially.
Sequential numbers can also be added for narrowed objects such as search results.
Sequential numbers can be added with the following search function.
- Statistics
- Keyword Search
- Feature Key Search
 Dongle License (HW Key)
Dongle License (HW Key) Feature Map
Feature Map Management and Operations of Feature Keys
Management and Operations of Feature Keys Sequence and Data Input and Output
Sequence and Data Input and Output GenBank EMBL Viewer
GenBank EMBL Viewer Sequence Viewer
Sequence Viewer Annotation Viewer
Annotation Viewer Circular Genome Viewer-Designer
Circular Genome Viewer-Designer Plasmid Map Viewer-Designer
Plasmid Map Viewer-Designer Trace Viewer - Editor
Trace Viewer - Editor Phylogenetic Tree Viewer
Phylogenetic Tree Viewer Feature Key Search
Feature Key Search Keyword Search
Keyword Search Pattern Search
Pattern Search Priming Site Search
Priming Site Search Batch Homology Search
Batch Homology Search Restriction Enzyme
Restriction Enzyme Primer Design
Primer Design PCR Reaction
PCR Reaction Ligation
Ligation Fragment Modification
Fragment Modification DNA Content Analysis
DNA Content Analysis Codon Analysis
Codon Analysis ORF Analysis
ORF Analysis Database Management
Database Management Multiple Circular Genome Map
Multiple Circular Genome Map Dot Plot Analysis
Dot Plot Analysis Venn Diagram Analysis
Venn Diagram Analysis Reverse Complement
Reverse Complement Settings
Settings