IMC i01K Draw Restriction Enzyme Map on Feature Map as a Lane
For the DNA sequence displayed in the current feature map, the recognition site of the selected restriction enzyme (s) is displayed as restriction enzyme map lane.
In addition to this, there is also a way to draw an independent restriction map different from the feature map.
In the latter case, you can continue to use linked functions such as cleavage of base sequence and gel electrophoresis.
In addition, there is a method to register and display the restriction enzyme recognition site as a feature on the feature lane of the main feature map.
Operation
- Load the nucleotide sequence to create the restriction enzyme map.
- The feature map is displayed.
- Click "Setting -> Layout Style" from the menu.
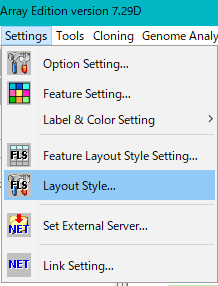
- The "Layout Style" dialog will be displayed.
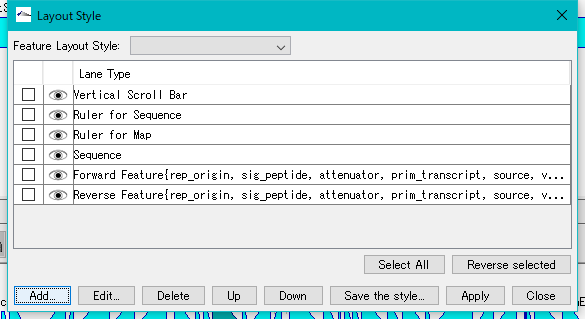
- Click "Add ...".
- The "Lane Style" dialog will be displayed.
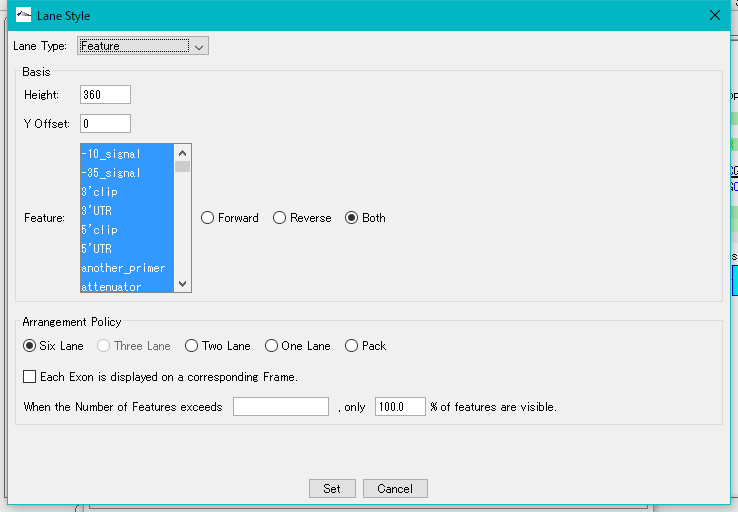
- Select "Enzyme Map" from the Lane Type pull-down menu.
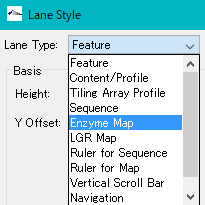
- The dialog changes to the screen for editing Enzyme Map.
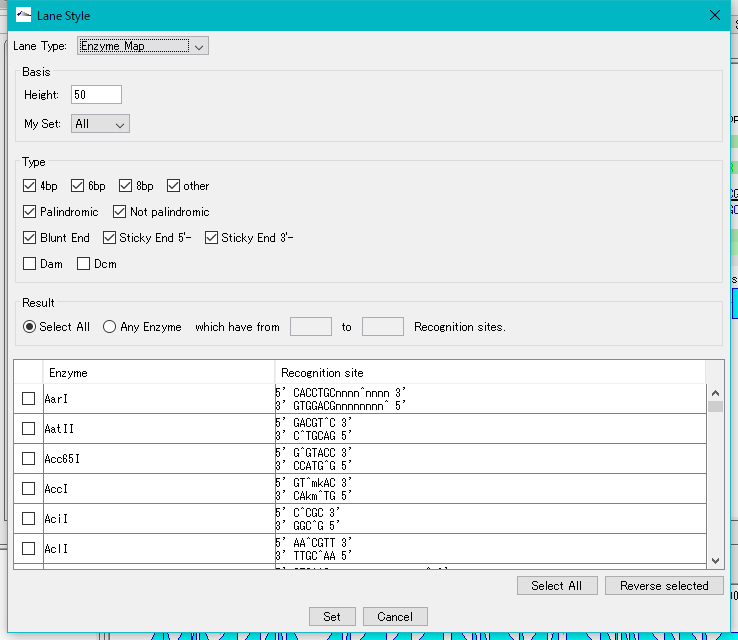
- From the Enzyme List, select the restriction enzyme to be displayed on the restriction enzyme map (multiple designation possible).
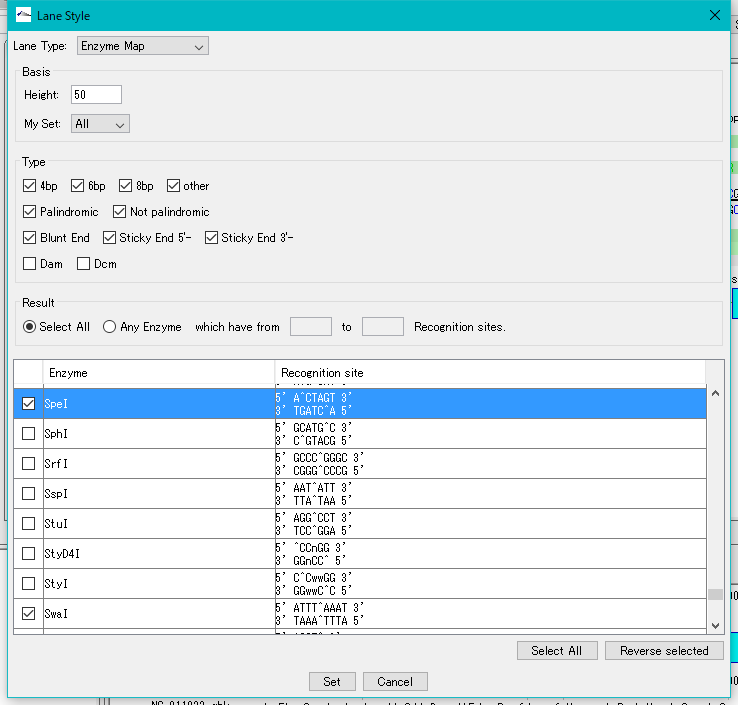
- Click the Set button.
- A row for Enzyme Map is added at the bottom of the style list of the Layout Style dialog.
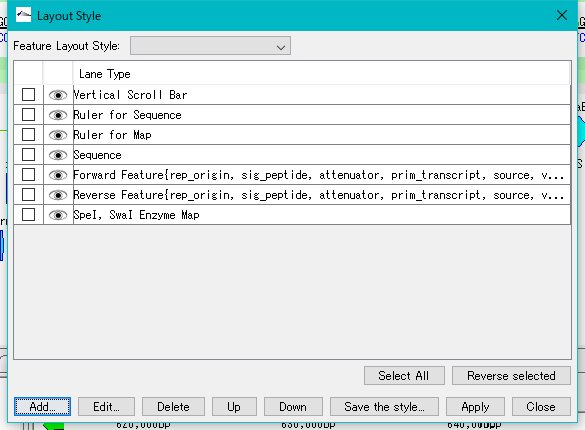
- Check the Enzyme Map line.
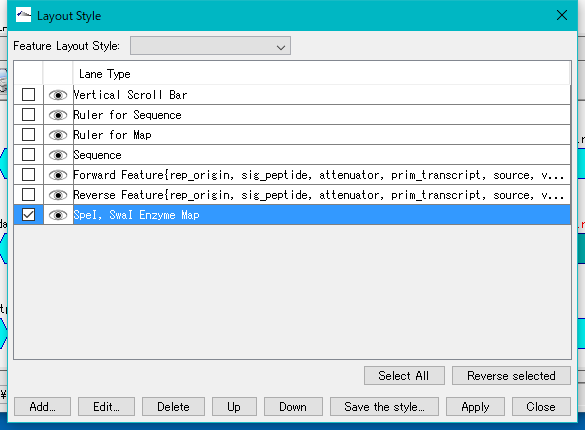
- Click UP several times and move the Enzyme Map line up.
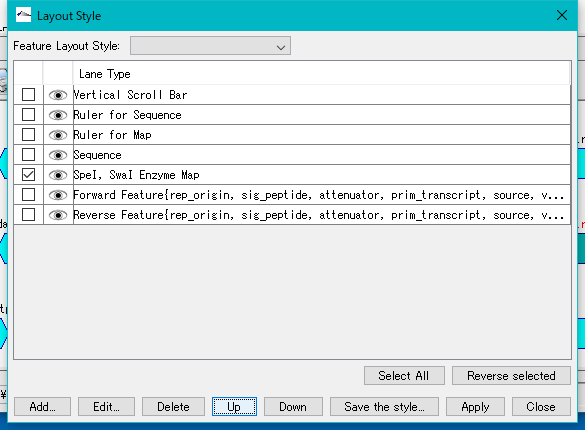
- Click the Apply button.
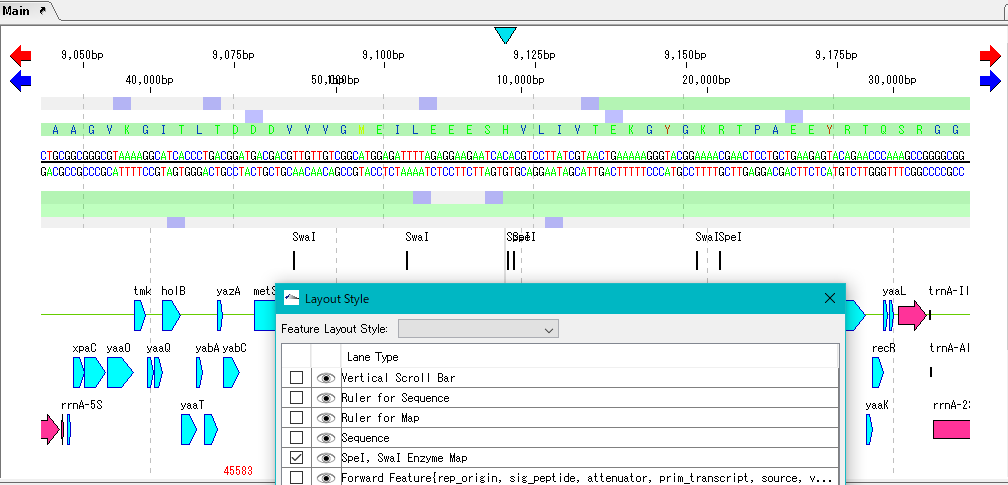
- The Layout Style dialog closes and the restriction enzyme map lane is displayed in the feature map.
 Dongle License (HW Key)
Dongle License (HW Key) Feature Map
Feature Map Management and Operations of Feature Keys
Management and Operations of Feature Keys Sequence and Data Input and Output
Sequence and Data Input and Output GenBank EMBL Viewer
GenBank EMBL Viewer Sequence Viewer
Sequence Viewer Annotation Viewer
Annotation Viewer Circular Genome Viewer-Designer
Circular Genome Viewer-Designer Plasmid Map Viewer-Designer
Plasmid Map Viewer-Designer Trace Viewer - Editor
Trace Viewer - Editor Phylogenetic Tree Viewer
Phylogenetic Tree Viewer Feature Key Search
Feature Key Search Keyword Search
Keyword Search Pattern Search
Pattern Search Priming Site Search
Priming Site Search Batch Homology Search
Batch Homology Search Restriction Enzyme
Restriction Enzyme Primer Design
Primer Design PCR Reaction
PCR Reaction Ligation
Ligation Fragment Modification
Fragment Modification DNA Content Analysis
DNA Content Analysis Codon Analysis
Codon Analysis ORF Analysis
ORF Analysis Database Management
Database Management Multiple Circular Genome Map
Multiple Circular Genome Map Dot Plot Analysis
Dot Plot Analysis Venn Diagram Analysis
Venn Diagram Analysis Reverse Complement
Reverse Complement Settings
Settings