- Map the amino acid sequence on the reference genome sequence and register it as a new feature
- Map the specified amino acid sequence (s) to the reference genome currently displayed as a feature map, and register the hit entry as a new feature (such as mRNA).
- The mapping uses the tBlastN algorithm.
How to operate amino acid sequence mapping
- Read the base sequence of the reference genome and display it on the feature map.
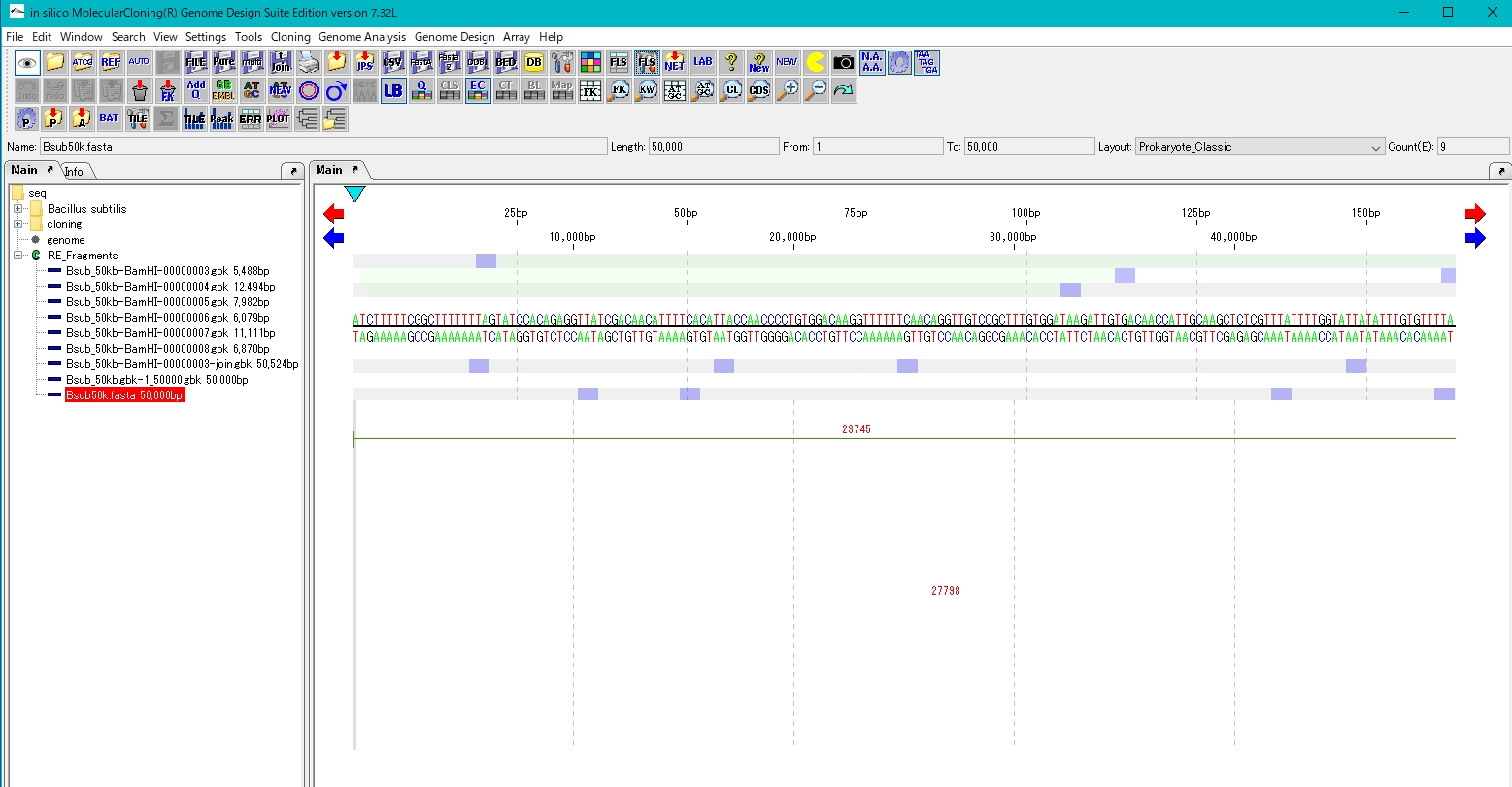
- Click Genome Analysis-> Mapping-> A.A. Sequence Mapping from the menu.
- A.A.Sequence Mapping execution dialog is displayed.
- Click Ref… of A.A.Sequence File (s) and specify the amino acid sequence to be mapped.
- The amino acid sequence that can be specified here is FastA format, GenPept format, etc. Files compressed with gz can also be specified.
- A dialog box for selecting an amino acid sequence appears.
- Specify the amino acid sequence file.
- Click the Run button.
- A confirmation message will be displayed.
- To change a parameter, click the Parameter button.
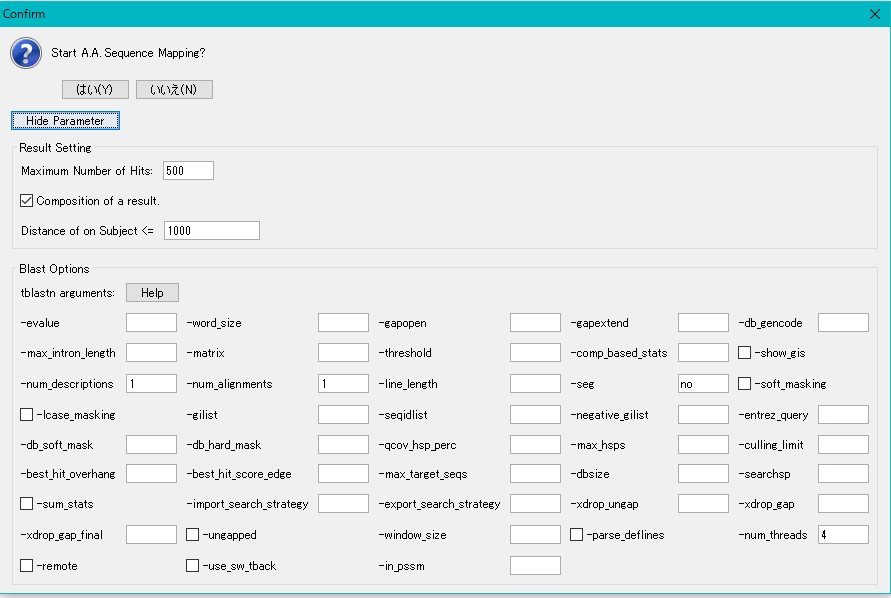
- This parameter is based on NCBI Blast parameter settings.
- Usually use the default parameter settings.
- Click the "Yes" button to start mapping.
- The execution time is proportional to the number and length of amino acid sequences and the base length of the reference genome base sequence.
- During execution, a message that indicates the processing progress is displayed.
- When the mapping process is completed, the result list dialog is displayed.
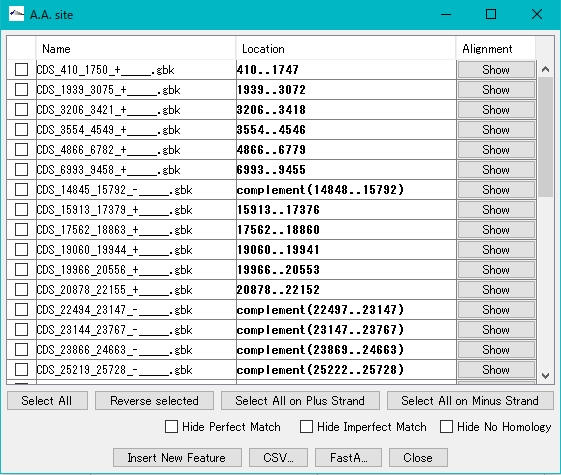
- Clicking on Alignment in each row will bring up a pairwise alignment dialog.
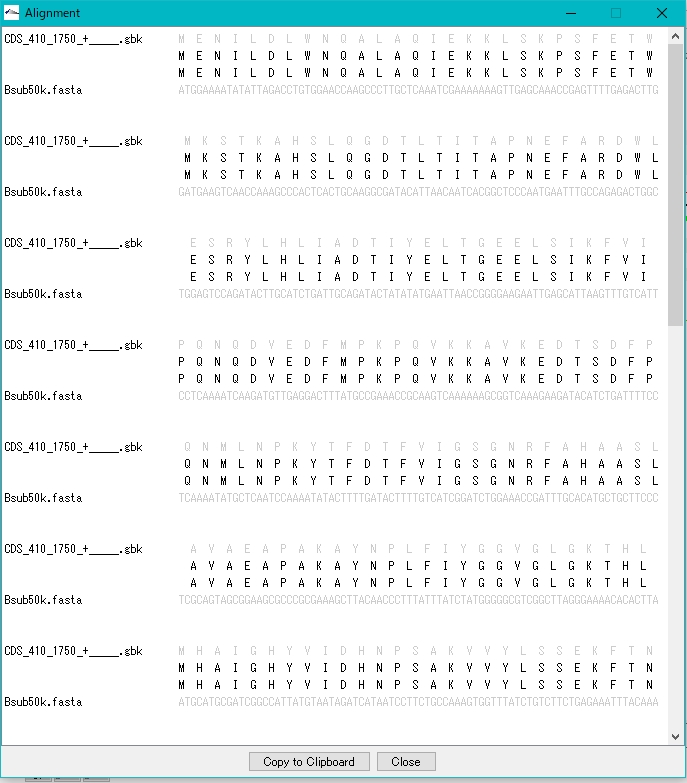
- Register a new feature using the result list dialog.
- Mapping results are classified into three types.
- When the input amino acid perfectly matches the reference genome sequence
- When the input amino acid incompletely matches the reference genome sequence
- When the input amino acid sequence does not match at all
- The former two types of results can be registered as new features on the reference genome. For example, a perfect match sequence can be registered as an mRNA feature, and an
- incomplete match sequence can be registered as a miscRNA feature. It is also possible to specify another feature key such as CDS.
- Click Insert Feature.
- A confirmation message will be displayed.
- Click “Yes”.
- The Feature Setting dialog is displayed.
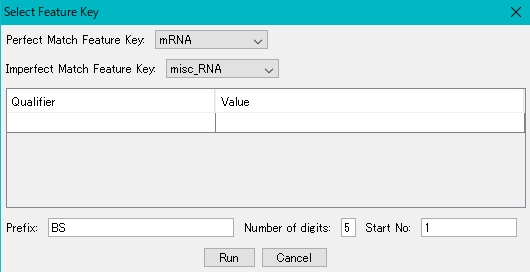
- The same value can be set to any qualifier of the newly registered feature. Also, you can register the prefix specified for Qualifier / locus_id and a serial number that has an arbitrary number of digits and starts with an arbitrary numerical value.

- In the case of eukaryotes with introns, exon and intron regions are identified and registered as features.
- You can limit the maximum base length of an intron.
- Click Run.
- When Feature registration is completed, a completion message is displayed.
- Click OK.
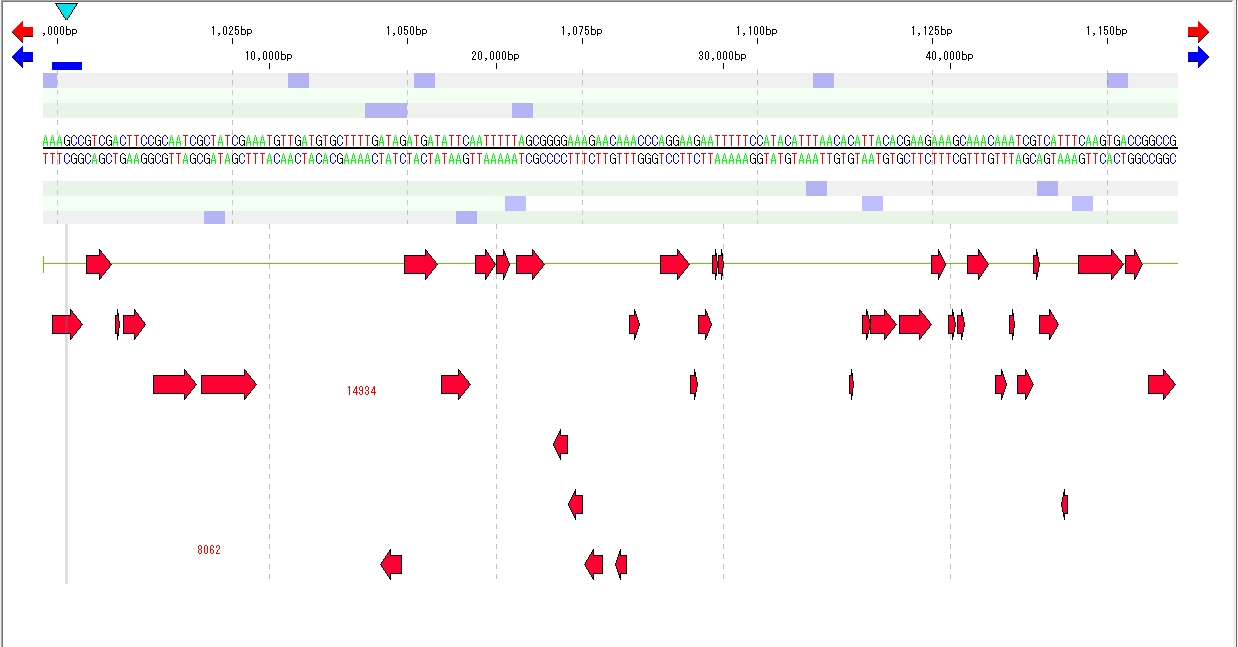
- The features registered in the main feature map are displayed.














 Dongle License (HW Key)
Dongle License (HW Key) Feature Map
Feature Map Management and Operations of Feature Keys
Management and Operations of Feature Keys Sequence and Data Input and Output
Sequence and Data Input and Output GenBank EMBL Viewer
GenBank EMBL Viewer Sequence Viewer
Sequence Viewer Annotation Viewer
Annotation Viewer Circular Genome Viewer-Designer
Circular Genome Viewer-Designer Plasmid Map Viewer-Designer
Plasmid Map Viewer-Designer Trace Viewer - Editor
Trace Viewer - Editor Phylogenetic Tree Viewer
Phylogenetic Tree Viewer Feature Key Search
Feature Key Search Keyword Search
Keyword Search Pattern Search
Pattern Search Priming Site Search
Priming Site Search Batch Homology Search
Batch Homology Search Restriction Enzyme
Restriction Enzyme Primer Design
Primer Design PCR Reaction
PCR Reaction Ligation
Ligation Fragment Modification
Fragment Modification DNA Content Analysis
DNA Content Analysis Codon Analysis
Codon Analysis ORF Analysis
ORF Analysis Database Management
Database Management Multiple Circular Genome Map
Multiple Circular Genome Map Dot Plot Analysis
Dot Plot Analysis Venn Diagram Analysis
Venn Diagram Analysis Reverse Complement
Reverse Complement Settings
Settings