IMC B2DA72 Extract ORF from Selection Area of Feature lane
Extract ORF candidates from the prokaryotic genome sequence and register them as new features.
The ORF candidate is judged by the length of the stop codon absence region.
You can set stop codon absence regions (ORF_long, ORF_Middle, ORF_Short) of three kinds of base length ranges.
ORF candidates can be changed to CDS by Translate function.
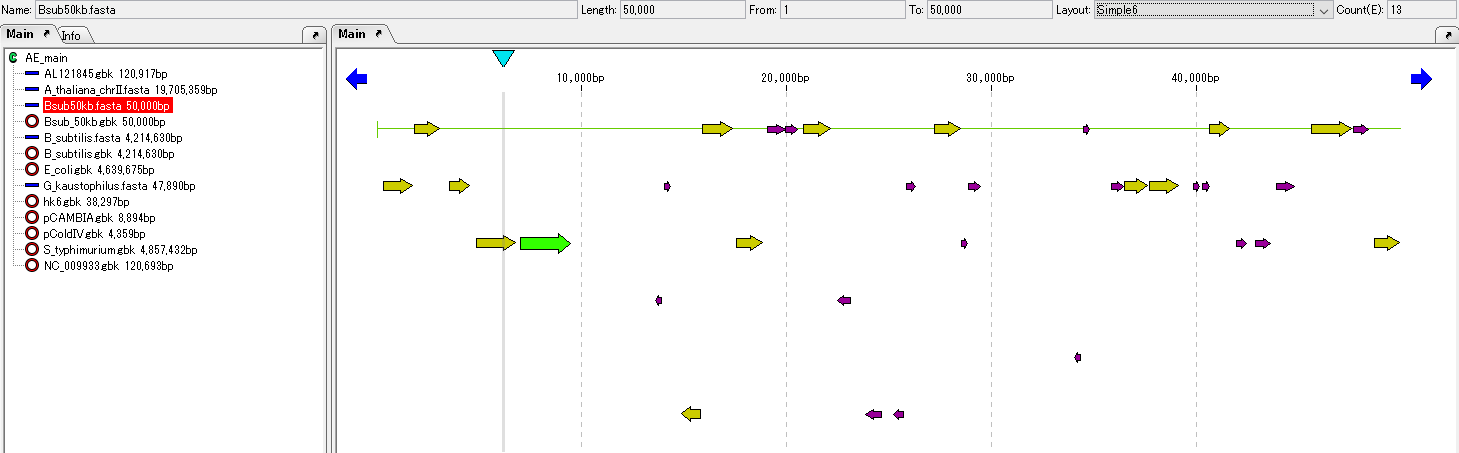
Operation
- Load the prokaryotic genomic sequence into the main current directory.
- As an example, we will use the sample data Bsub 50 kb.fasta.
- When loading, it usually becomes the current arrangement and it is displayed in the main feature map.
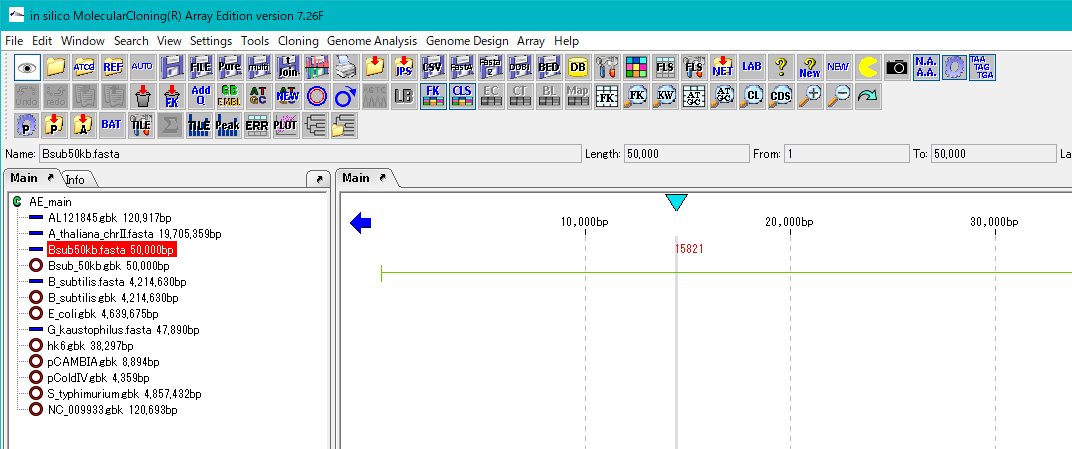
- If it does not appear in the main feature map, click this array on the main directory.
- Click "Genome Analysis -> ORF -> ORF Extraction" from the menu.
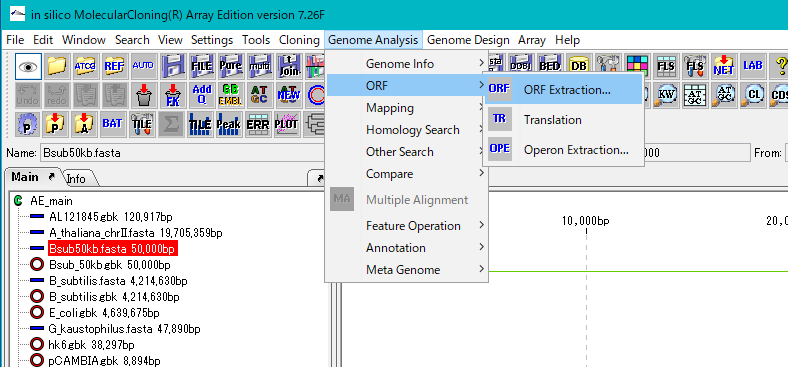
- The "Range Setting" dialog will be displayed.
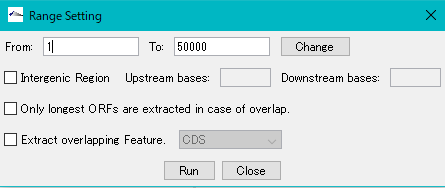
- Check "Only Longest ORFs are Extracted in case of Overlap" checkbox.
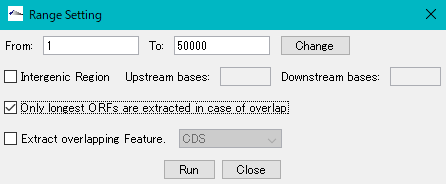
- Click "Run".
- The "Stard ORF Search?" Confirmation message is displayed.
- Click "Yes (Y)".
- When the ORF extraction is completed, the "ORF Site" dialog will be displayed.
- The number of ORF candidate detections by ORF type is displayed in the list on the left.
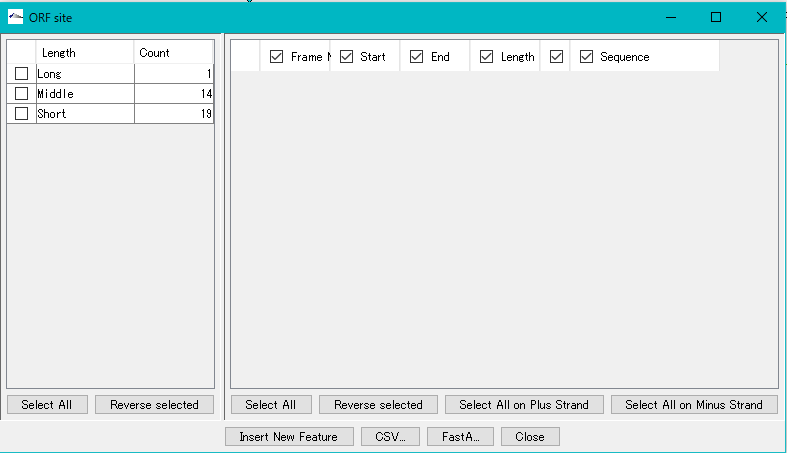
- Check each check box in the list on the left.
- Alternatively, click "Select All" in the lower left.
- The frame No., start base position, end base position, base length, strand and base of each ORF candidate are displayed in the list on the right.
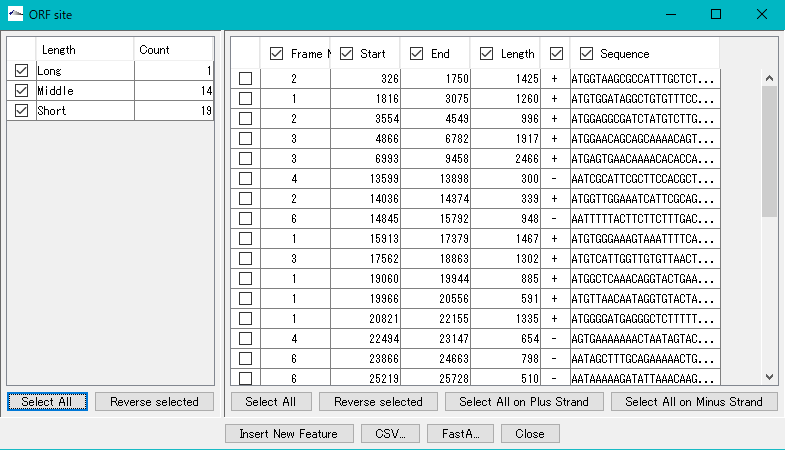
- To register all extracted ORF candidates as features, click "Select All" in the lower right.
- All ORF candidates are checked.
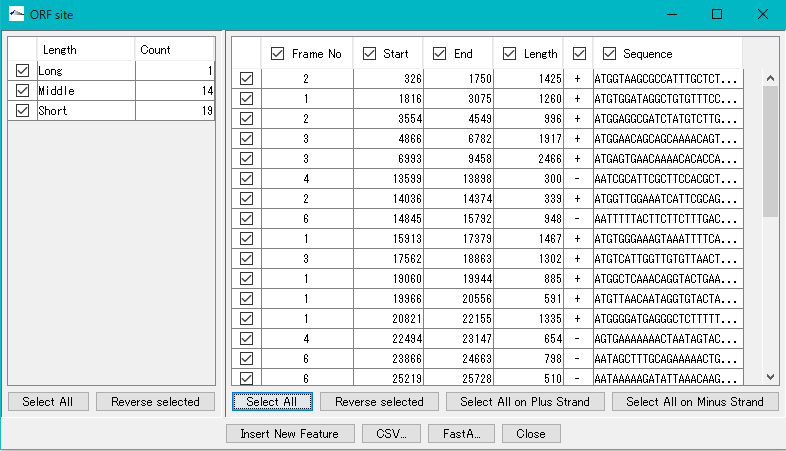
- Click "Insert New Feature".
- The "Insert New Feature?" Confirmation message will be displayed.
- Click "Yes (Y)".
- A "Completed !!!" message will be displayed.
- Click "OK".
- If you do not close the "ORF Site" dialog, you can use it to check the position of each ORF candidate.
- Click one of the ORF candidates in the right list.
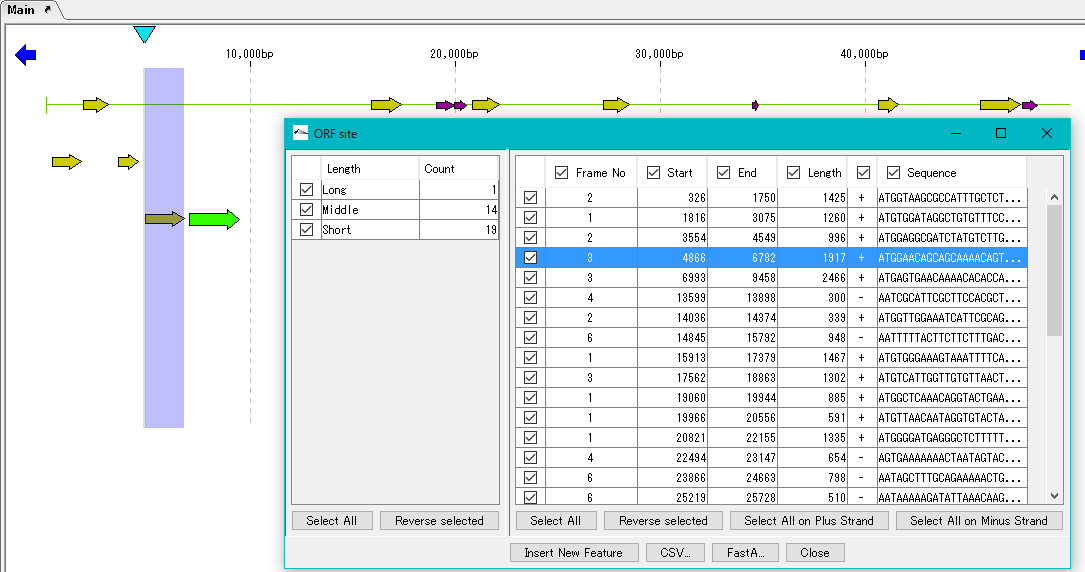
- The position of the corresponding feature in the main feature map is highlighted.
- Click "Close" in "ORF Site" dialog.
- The "ORF Site" dialog closes.
- Feature of registered ORF candidate is displayed on the feature lane of the main feature map.

After this, proceed to "Change the candidate's feature key to CDS and translate amino acids".
 Dongle License (HW Key)
Dongle License (HW Key) Feature Map
Feature Map Management and Operations of Feature Keys
Management and Operations of Feature Keys Sequence and Data Input and Output
Sequence and Data Input and Output GenBank EMBL Viewer
GenBank EMBL Viewer Sequence Viewer
Sequence Viewer Annotation Viewer
Annotation Viewer Circular Genome Viewer-Designer
Circular Genome Viewer-Designer Plasmid Map Viewer-Designer
Plasmid Map Viewer-Designer Trace Viewer - Editor
Trace Viewer - Editor Phylogenetic Tree Viewer
Phylogenetic Tree Viewer Feature Key Search
Feature Key Search Keyword Search
Keyword Search Pattern Search
Pattern Search Priming Site Search
Priming Site Search Batch Homology Search
Batch Homology Search Restriction Enzyme
Restriction Enzyme Primer Design
Primer Design PCR Reaction
PCR Reaction Ligation
Ligation Fragment Modification
Fragment Modification DNA Content Analysis
DNA Content Analysis Codon Analysis
Codon Analysis ORF Analysis
ORF Analysis Database Management
Database Management Multiple Circular Genome Map
Multiple Circular Genome Map Dot Plot Analysis
Dot Plot Analysis Venn Diagram Analysis
Venn Diagram Analysis Reverse Complement
Reverse Complement Settings
Settings