IMC G2B Setting Dialog for Sequence Lane
In the Setting dialog, Sequence Lane tab pane, change the settings such as the color and size of bases and amino acid characters in the feature lane of the main feature map.
The sequence (sequence) lane consists of 6 rows, each of which has a base sequence of the genome (regular chain, opposite strand) line by line as shown below, and in the coding region, the amino acid sequence is a forward chain and a reverse sequence each 3 frames It is.

Feature setting Tab Pane
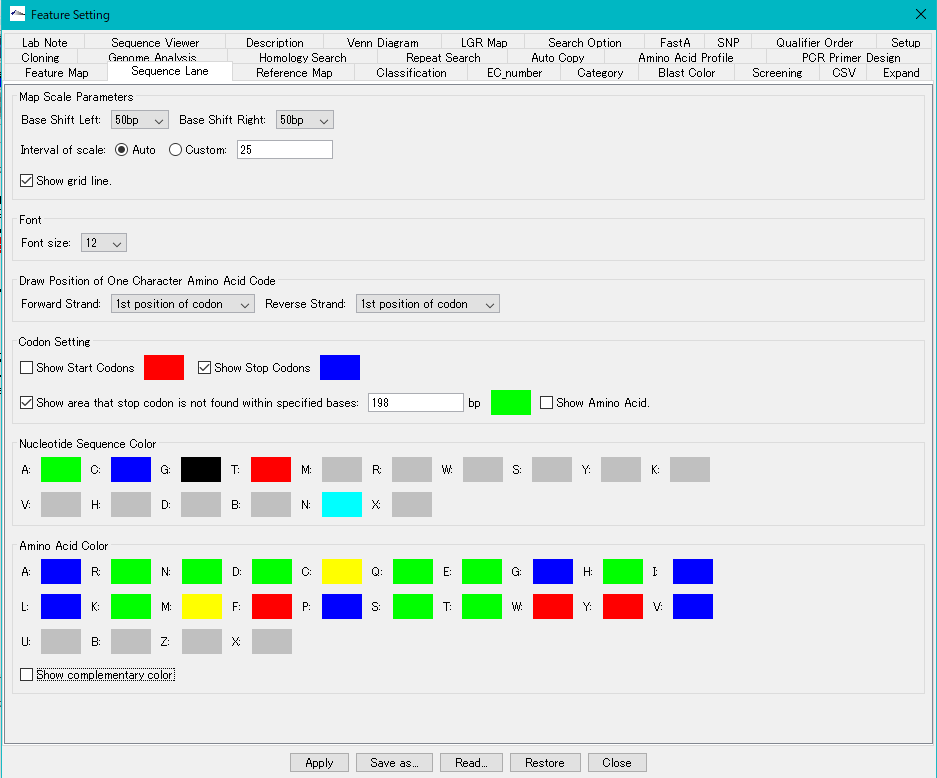
Map Scale Parameters
Base Shift Left: From the pull-down menu, specify the number of scroll bases per scroll of the left scroll button or the scroll width with the screen width set to 1.
Base Shift Right: From the pull-down menu, specify the scroll base number per one click of the right scroll button or the scroll width with the screen width set to 1.
Interval of Scale: Specify the scale interval of the scale lane. You can select from Auto and Custom, and if you select Custom, you can specify the scale interval in base units.
Show Grid Line check box: When checked, vertical grid lines are displayed at tick marks on the array lane.
Font
Font Size: You can select the font size of the array character from the pull-down menu.
Draw Position of One Character Amino Acid Code
Forward Strand: On the amino acid display area of the normal chain, specify the position in the codon where the one-letter amino acid code is displayed from the pull-down menu.
Reverse Strand: On the amino acid display region of the opposite chain, specify the position in the codon where the one-letter amino acid code is displayed from the pull-down menu.
Codon Setting
Show Start Codon check box: When checked, the position of the start codon candidate is displayed as a rectangle in the amino acid display area. The start codon is referenced from the applied Genetic Table. Click the color box to change the display color. In addition, color, transparency is different from the color specified in the color box.
Show Stop Codon check box: When checked, the position of the stop codon candidate is displayed as a rectangle in the amino acid display area. The stop codon is referenced from the applied Genetic Table. Click the color box to change the display color. In addition, color, transparency is different from the color specified in the color box.
Show Area that Stop Codon is not Found within Specified Bases check box: Fills the area with the color specified in the color box when there is no stop codon in the area that is equal to or more than the number of bases specified in the input field.
Show Amino Acid check box: When checked, the amino acid sequence when translating this area is displayed.
Nucleotide Sequence Color color box: Specify the base letter color of the sequence lane for each nucleotide code.
Amino Acid Color color box: For each amino acid code, specify the amino acid character color to be displayed in the amino acid display area of the sequence lane.
Show Complementary Color check box: The background of the amino acid code to be checked is displayed in the specified color, and the characters themselves are displayed in complementary colors.
 Dongle License (HW Key)
Dongle License (HW Key) Feature Map
Feature Map Management and Operations of Feature Keys
Management and Operations of Feature Keys Sequence and Data Input and Output
Sequence and Data Input and Output GenBank EMBL Viewer
GenBank EMBL Viewer Sequence Viewer
Sequence Viewer Annotation Viewer
Annotation Viewer Circular Genome Viewer-Designer
Circular Genome Viewer-Designer Plasmid Map Viewer-Designer
Plasmid Map Viewer-Designer Trace Viewer - Editor
Trace Viewer - Editor Phylogenetic Tree Viewer
Phylogenetic Tree Viewer Feature Key Search
Feature Key Search Keyword Search
Keyword Search Pattern Search
Pattern Search Priming Site Search
Priming Site Search Batch Homology Search
Batch Homology Search Restriction Enzyme
Restriction Enzyme Primer Design
Primer Design PCR Reaction
PCR Reaction Ligation
Ligation Fragment Modification
Fragment Modification DNA Content Analysis
DNA Content Analysis Codon Analysis
Codon Analysis ORF Analysis
ORF Analysis Database Management
Database Management Multiple Circular Genome Map
Multiple Circular Genome Map Dot Plot Analysis
Dot Plot Analysis Venn Diagram Analysis
Venn Diagram Analysis Reverse Complement
Reverse Complement Settings
Settings