Function Menu
 License Activation
License Activation Function Overviews and Basic Operation
Function Overviews and Basic Operation Work Bench
Work Bench Feature Map
Feature Map- Feature Key and Feature
 Management and Operations of Feature Keys
Management and Operations of Feature Keys- Attributes of Feature Keys
- IMC Original Set Feature Keys
- Types and Roles of Qualifiers
- IMC Original Set Qualifiers
- View and Edit Qualifiers
- Feature Position on Genome Sequence
- Feature Fragmentation
- Feature Synthesis
- Feature Fusion
- Feature Operators
- Link and Refer on Feature
- Feature Mapping
- Register, Edit, Delete Feature
- Feature Appearance
- Join and Delete Position
- Feature Categorization and Presentation
- Import Feature
- Export Feature
- Numbering of Feature
 Sequence and Data Input and Output
Sequence and Data Input and Output- Genome / Sequence Viewer / Editor
 GenBank EMBL Viewer
GenBank EMBL Viewer Sequence Viewer
Sequence Viewer Annotation Viewer
Annotation Viewer- Multiple Genome Viewer (Linear Map)
 Circular Genome Viewer-Designer
Circular Genome Viewer-Designer Plasmid Map Viewer-Designer
Plasmid Map Viewer-Designer Trace Viewer - Editor
Trace Viewer - Editor- Labeling and Coloring
- Description Window
- Amino Acid Sequence Profile Viewer
- Multiple Alignment Viewer
 Phylogenetic Tree Viewer
Phylogenetic Tree Viewer- GT Alignment Viewer
- Restriction Enzyme Map Window
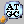 Search Sequence and Annotation
Search Sequence and Annotation- Cloning
- Sequencing
- Gene and Genome Sequence Analysis
- Genome Annotation
 Genome Comparison
Genome Comparison- Multiple Aligment
 Phylogenetic Tree
Phylogenetic Tree- Multiple Linear Genome Map
- Gene Cluster Alignment
 Multiple Circular Genome Map
Multiple Circular Genome Map Dot Plot Analysis
Dot Plot Analysis Venn Diagram Analysis
Venn Diagram Analysis- Core Genome Analysis
- Global Genome Rearrangement Analysis
- Local Genome Rearrangement Analysis
- Mutation Analysis
- Enzyme Alignment by EC Number
- Unique Region Analysis
- Genome Mapping
- Expression Analysis
- Metabolome Analysis
- Genome Design
 Settings
Settings- Tools
- Window and Dialog Description
- What is "Do It Yourself" Genome Analysis Software?
- Functions and Operations of GenomeTraveler
You are here: Home  Genome Comparison
Genome Comparison  Gene Cluster Alignment
Gene Cluster Alignment  IMC O10 How to Draw Gene Cluster Alignment
IMC O10 How to Draw Gene Cluster Alignment
 Genome Comparison
Genome Comparison  Gene Cluster Alignment
Gene Cluster Alignment  IMC O10 How to Draw Gene Cluster Alignment
IMC O10 How to Draw Gene Cluster AlignmentIMC O10 How to Draw Gene Cluster Alignment
| | |
Print | Hits: 5520
How to perform gene cluster alignment between closely related genomes.
- This function can be used with the following editions.
- IMC GE
 AE
AE DS
DS
Operation
- Load the annotated (gene-identified) hall genome nucleotide sequence that you want to compare to the current reference directory.
- Right-click on the genes of interest (CDS) of one genome of the reference feature map.
- Select "Gene Cluster Alignment" from the displayed menu.
- The execution dialog is displayed. Click RUN.
- An execution confirmation message is displayed.
- Click "Yes".
- When execution is started and the process ends, the Binary Best Hit List window will be displayed, and homologous genes around the reference gene on the reference feature map will be joined by bands.
- Even if you scroll through one genome, the band moves accordingly, but the topology does not change.
- Even if you zoom, it is the same.
| Category: Genome Comparison / Gene Cluster Alignment
Related Articles
- 2018-11-25 - IMC F08A Launch Phylogenetic Tree Viewer from Multiple Alignment Viewer
- 2018-08-14 - IMC Ver. 7.24 is released
- 2018-08-22 - IMC Ver. 7.25 is released
- 2018-08-28 - IMC Version 7.26 is released
- 2018-12-07 - IMC S411 Execute OGAB Fragmentation Designer
- 2018-12-29 - GT F31A Launch GT Alignment Viewer from Analysis Node (BAM)
- 2018-10-31 - IMC Ver. 7.27 is released
- 2018-09-24 - IMC E04G Pairwise Alignment between Query and Hit Entry
- 2018-11-22 - IMC O10A Operation of Gene Cluster Alignement
- 2018-11-25 - IMC F09A Launch Multiple Alignment Viewer from Homology Search Result
Site Seal
Full-Text Search the Site
Recent Updates
- IMC Version 8.135 Release Note
- IMC Version 8.133 Release Note
- IMC E03A Overviews on Pattern Search
- IMC i02P Primer Modification
- IMC Version 8.130 Release Note
- IMC Version 8.129 Release Note
- IMC Version 8.132 Release Note
- IMC Version 8.128 Release Note
- IMC Version 8.127 Release Note
- IMC Version 8.126 Release Note
- IMC O04 Venn Diagram Overviews
- IMC L01B Load Related Genome to Reference Map, Auto Generation of DB
- IMC i03A execute PCR
- IMC i06E Extrace Band from Gel
- IMC i03B Register Priming Site as a Feature
- IMC i02J Primer Desing for In-Fusion PCR
- IMC Version 8.125 Release Note
- IMC Version 8.123 Release Note
- IMC Version 8.111 Release Note
- IMC N01A2: about Genome Scale Model in Excel Format
Popular Tags in the Site
trouble
5
SNP
6
Qualifier
10
navigation
4
Mutation
5
Local Genome Rearrangement Map
6
Linear Map
9
License
5
Lane
9
Genome Design
7
Gene Cluster Alignment
8
Frame
6
Feature Map
6
Feature Layout Style
5
feature key
5
Feature
6
download
4
Codon
4
BLAST
11
Activation
4
 Dongle License (HW Key)
Dongle License (HW Key) Feature Key Search
Feature Key Search Keyword Search
Keyword Search Pattern Search
Pattern Search Priming Site Search
Priming Site Search Batch Homology Search
Batch Homology Search Restriction Enzyme
Restriction Enzyme Primer Design
Primer Design PCR Reaction
PCR Reaction Ligation
Ligation Fragment Modification
Fragment Modification DNA Content Analysis
DNA Content Analysis Codon Analysis
Codon Analysis ORF Analysis
ORF Analysis Database Management
Database Management Reverse Complement
Reverse Complement