IMC i01L Linearize Circular Vector by Restriction Enzyme
Linearize circular vector with restriction enzyme
Operation
- Load the DNA sequence of a circular vector into the current main directory.
- Load the sample sequence "pColdV.gbk" as an example.
- The sequence loaded in the main feature map is displayed.
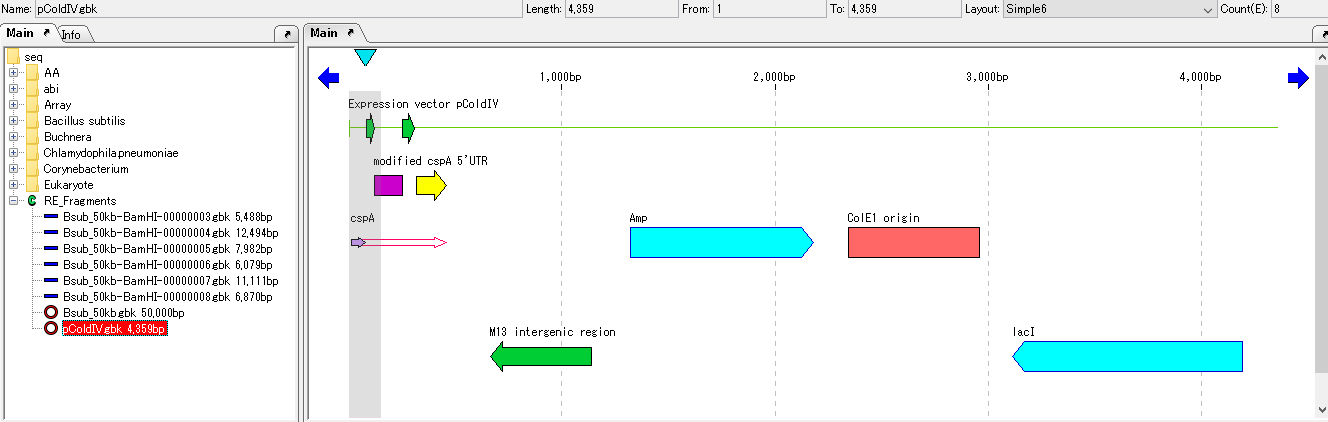
- Find a restriction enzyme that cleaves this vector in only one place.
- Click the "Cloning -> Restriction Enzyme -> RE Registration & Editing ..." button from the menu.
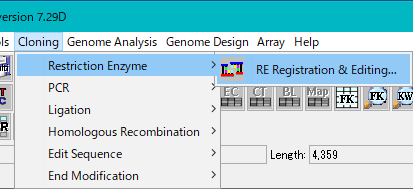
- The "Enzyme Selection" dialog is displayed.
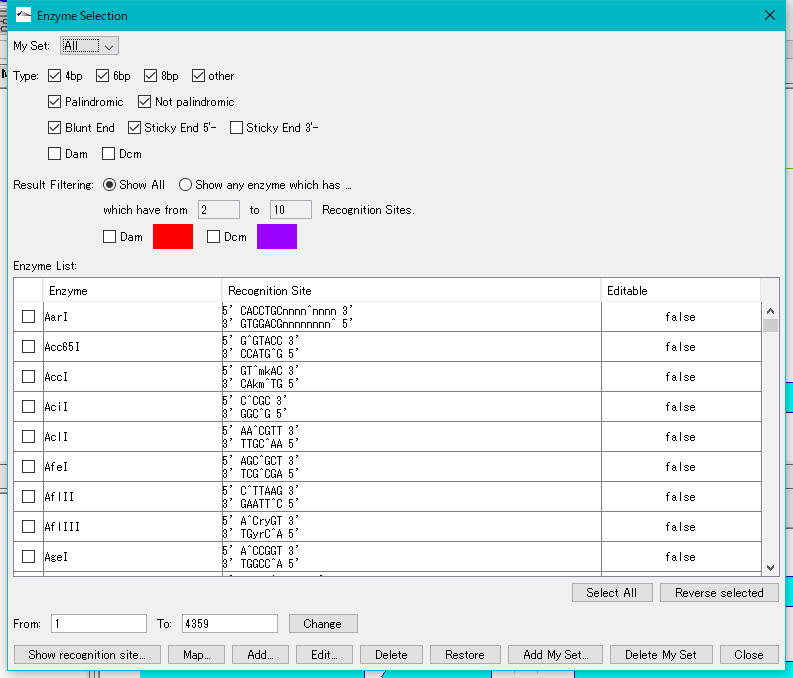
- In the "Select Enzyme" section, in the "Result" field, check "Any Enzyme".
- Enter 1 in the "which have from" and "To" fields.
- Click "Select All". All restriction enzymes are checked.
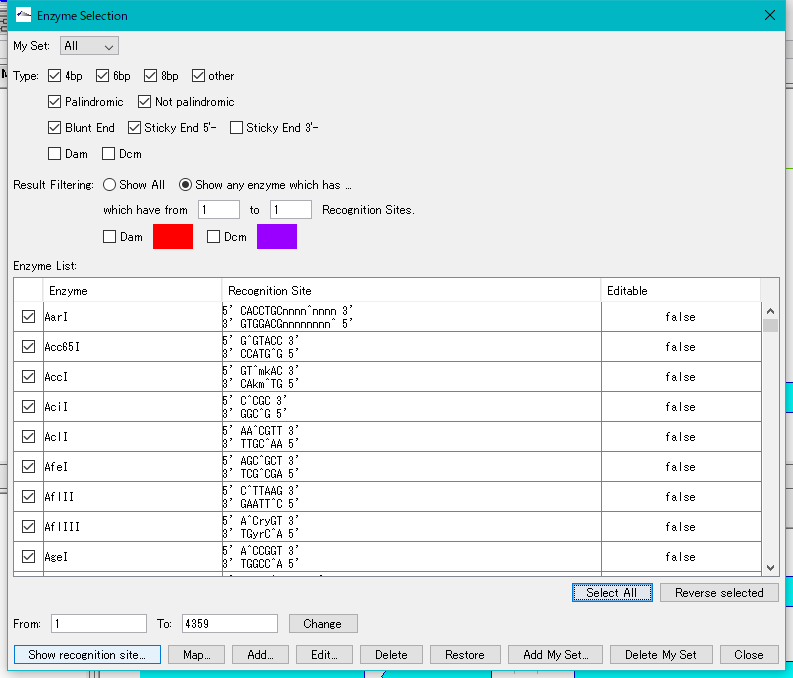
- Click "Show Recognition Site ...".
- The "Recognition Site" dialog is displayed.
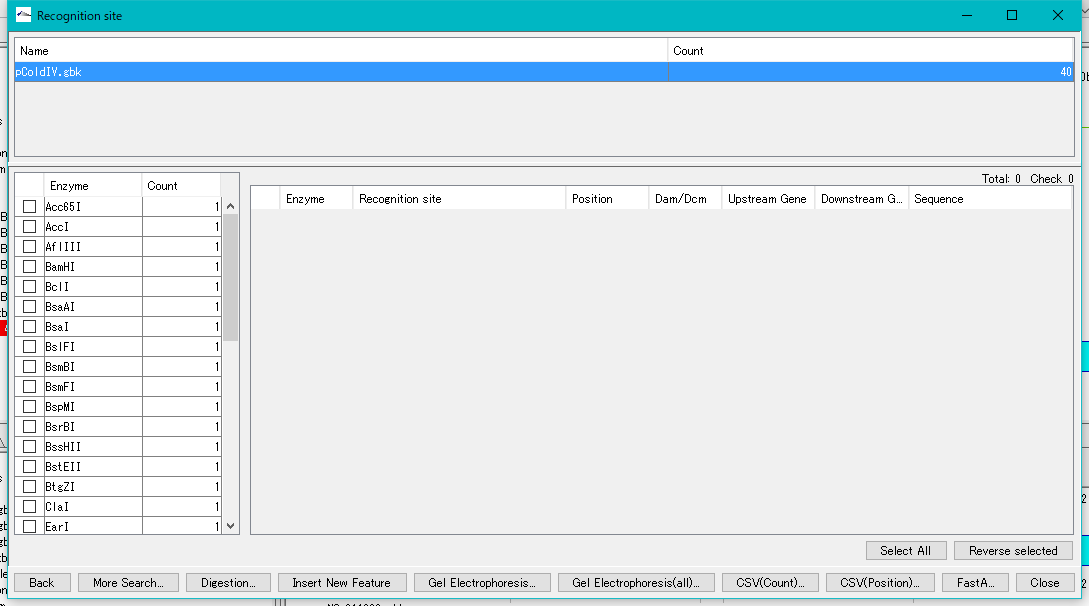
- For example, you know that BamHI disconnects MCS in only one place, so check BamHI.
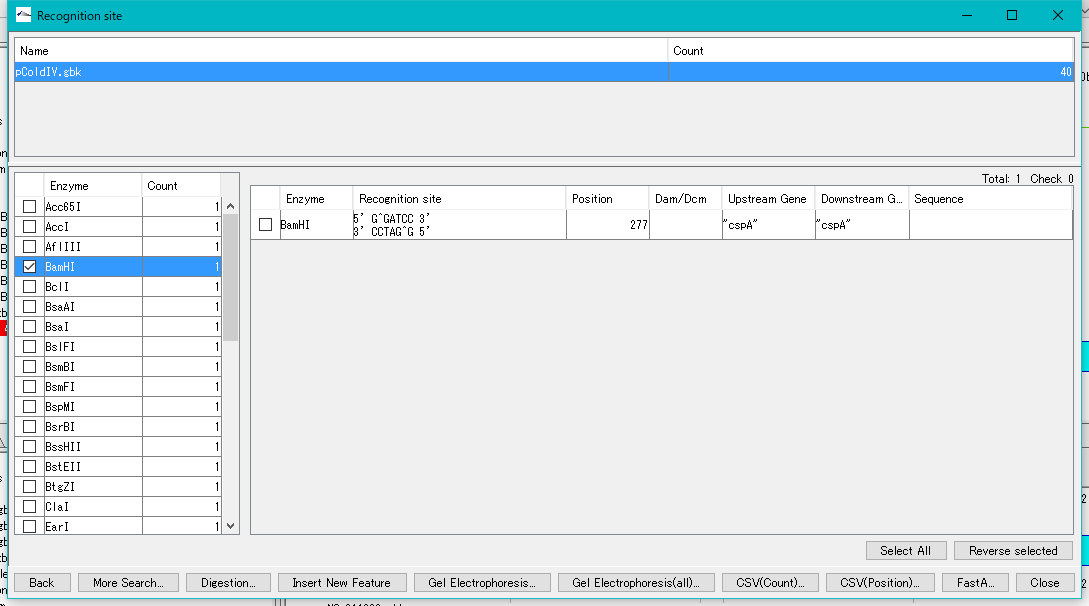
- The recognition site is displayed in the right pane.
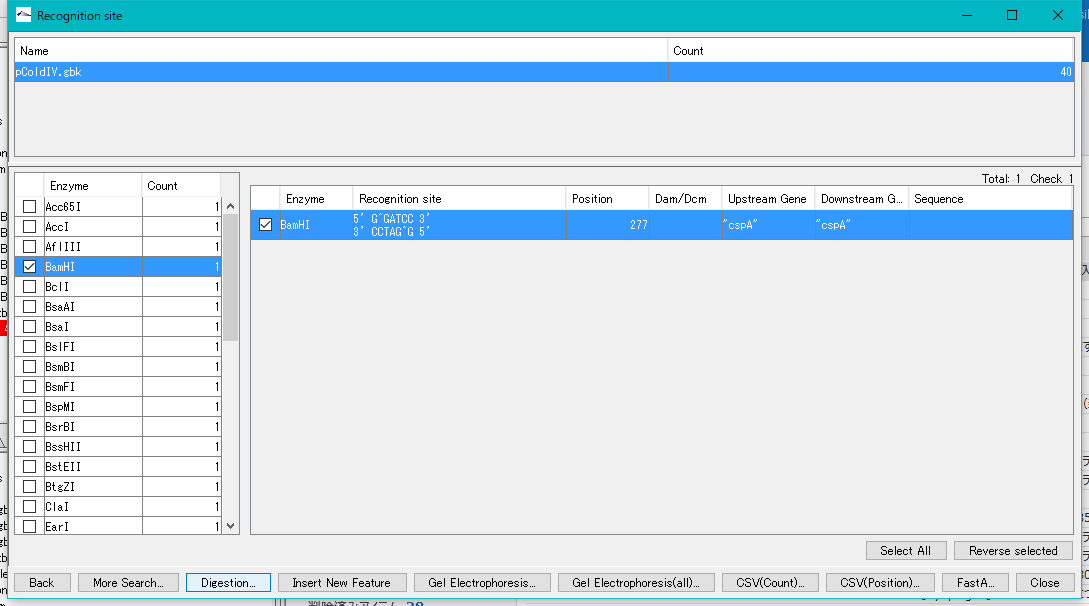
- Check the BamHI in the right pane and click the "Digestion" button.
- A confirmation message will be displayed.
- Click "Yes (Y)".
- The "Digestion list" window will be displayed.
- Only one digested fragment is listed.
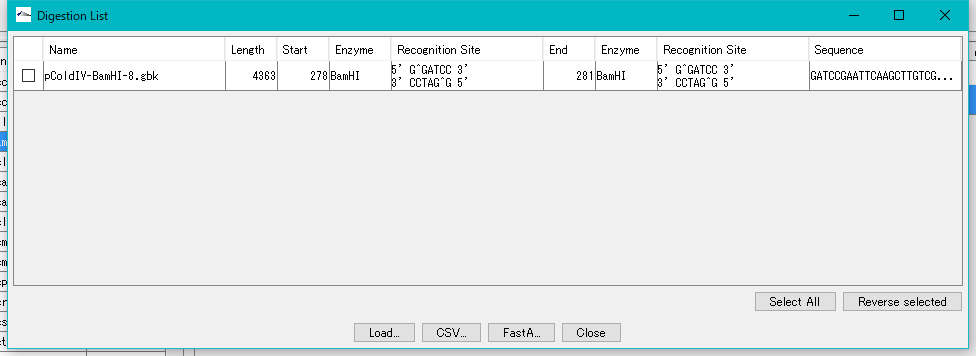
- Check the fragment and click the "Load" button.
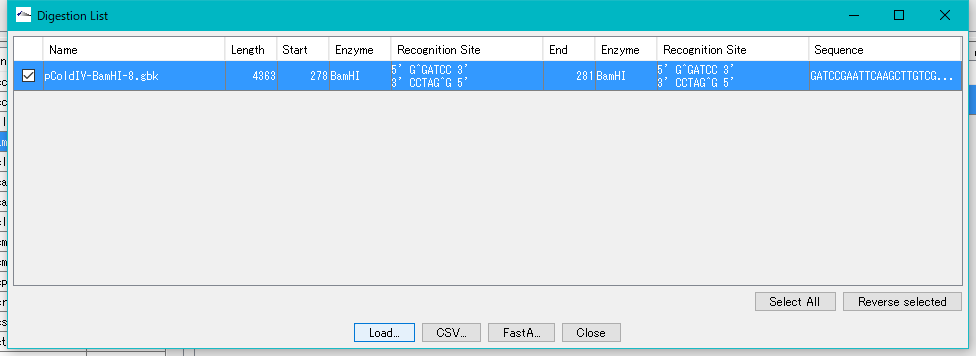
- A "Completed !!" completion message is displayed.
- Click "OK".
- The vector sequence opened with the main feature map is loaded.
- Linearized DNA has end points for horizontal scrolling.
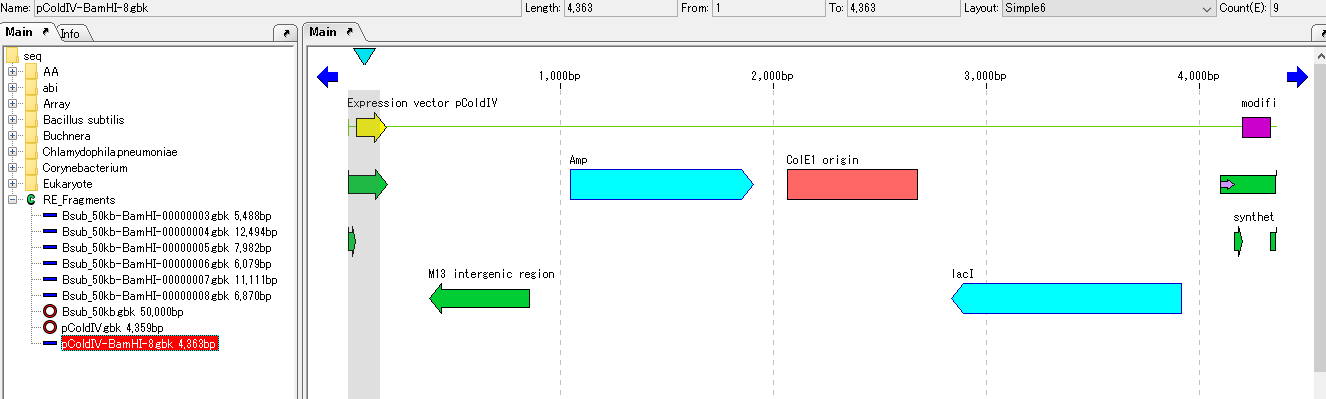
- Click "Cloning -> Ligation -> Simple Ligation ..." from the menu.
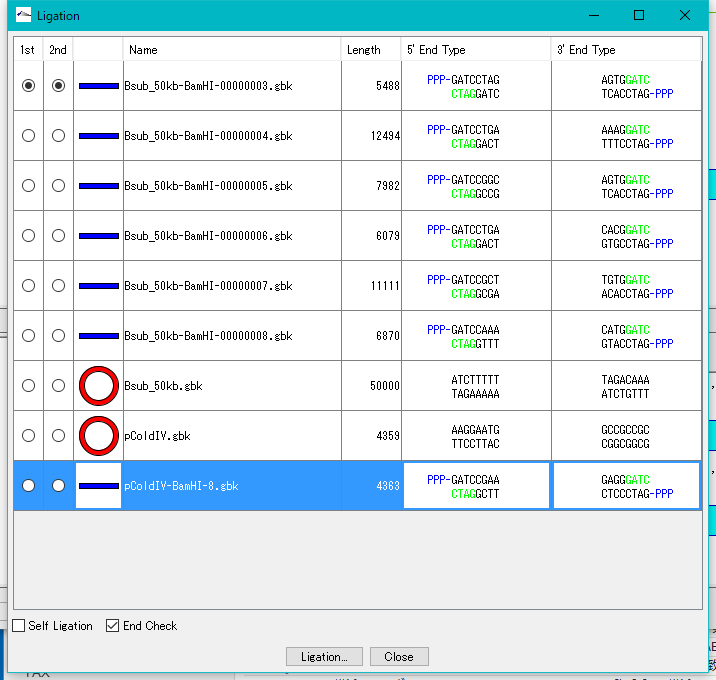
- The Ligation dialog will be displayed.
- There is a vector opened at the bottom of the list, and the shape of both ends is displayed.
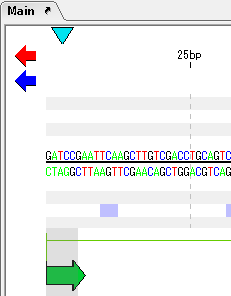
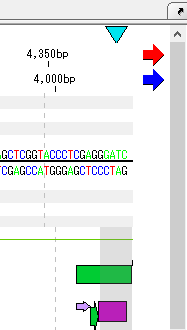
- The end shape can also be seen in the sequence lane of the main feature map.
- Scroll horizontally until scrolling stops.
 Dongle License (HW Key)
Dongle License (HW Key) Feature Map
Feature Map Management and Operations of Feature Keys
Management and Operations of Feature Keys Sequence and Data Input and Output
Sequence and Data Input and Output GenBank EMBL Viewer
GenBank EMBL Viewer Sequence Viewer
Sequence Viewer Annotation Viewer
Annotation Viewer Circular Genome Viewer-Designer
Circular Genome Viewer-Designer Plasmid Map Viewer-Designer
Plasmid Map Viewer-Designer Trace Viewer - Editor
Trace Viewer - Editor Phylogenetic Tree Viewer
Phylogenetic Tree Viewer Feature Key Search
Feature Key Search Keyword Search
Keyword Search Pattern Search
Pattern Search Priming Site Search
Priming Site Search Batch Homology Search
Batch Homology Search Restriction Enzyme
Restriction Enzyme Primer Design
Primer Design PCR Reaction
PCR Reaction Ligation
Ligation Fragment Modification
Fragment Modification DNA Content Analysis
DNA Content Analysis Codon Analysis
Codon Analysis ORF Analysis
ORF Analysis Database Management
Database Management Multiple Circular Genome Map
Multiple Circular Genome Map Dot Plot Analysis
Dot Plot Analysis Venn Diagram Analysis
Venn Diagram Analysis Reverse Complement
Reverse Complement Settings
Settings